COVID-19 diagnostic study of choice
For COVID-19 frequently asked inpatient questions, click here
For COVID-19 frequently asked outpatient questions, click here
|
COVID-19 Microchapters |
|
Diagnosis |
|---|
|
Treatment |
|
Case Studies |
|
COVID-19 diagnostic study of choice On the Web |
|
American Roentgen Ray Society Images of COVID-19 diagnostic study of choice |
|
Risk calculators and risk factors for COVID-19 diagnostic study of choice |
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [1] ; Associate Editor(s)-in-Chief: Sabawoon Mirwais, M.B.B.S, M.D.[2] Mydah Sajid, MD[3]
Overview
The diagnostic study of choice for COVID-19 illness is the Nucleic acid amplification test (NAAT) by a reverse transcriptase-polymerase chain reaction (RT-PCR). The RT-PCR is performed in symptomatic patients and individuals with high-risk exposure. There is no need to perform additional tests in situations with positive NAAT. The diagnostic criteria for suspected and confirmed cases of coronavirus disease 2019 (COVID-19) are also tabulated in the section below.
Diagnostic Study of Choice
Study of Choice
- The Nucleic acid amplification test by the reverse transcriptase-polymerase chain reaction (RT-PCR) is the gold standard test for detecting coronavirus illness. These tests are highly specific for detecting coronavirus illness.[1]
Diagnostic Results
- The NAAT detects viral gene sequence specific for coronavirus. The genes detected by NAAT include nucleocapsid(N), envelope(E), spike(S), and RNA dependent RNA polymerase (RdRP) in the open reading frame. [2]
- The turnaround time for these tests is variable depending upon the specific testing kit and laboratory performing it.[3]
- The diagnosis of coronavirus illness requires detection of at least two genes specific for SAR Cov-2 by RT-PCR. In the United States, the Center for disease control and prevention recommends testing for two nucleocapsid proteins N1 and N2.[4]
- World Health Organization recommends two-step gene testing by NAAT for diagnosis of coronavirus illness. The first step is screening for coronavirus illness with Envelope(E) gene assay followed by a confirmatory test with RNA dependent RNA polymerase (RdRP) gene assay.[5]
- The RdRp/Hel gene assay has the highest sensitivity and specificity for the diagnosis of coronavirus illness in comparison with spike and nucleocapsid genes. It has the lowest limit of detection in vitro.[6]
- The NAAT can have false-negative results. The false-negative results can depend upon a number of factors related to sample, course of illness, genetic mutations in the virus.[7] An adequate sample should be taken with proper technique and transported and stored at an optimum temperature from the testing area to laboratory.
- The sampling time on the day of illness is important as the viral load is low in the early and convalescence stages of infection. The RT-PCR is positive one to two days before the onset of symptoms till 7 to 12 days in moderate cases and up to two weeks in severe patients.[7] A study was done in a hospital in China where 51 coronavirus suspected patients had RT-PCR tests and chest CT scans. 36 patients had positive first RT-PCR tests. 15 patients had negative tests with CT scan findings were highly suggestive of viral pneumonia. Serial testing by RT-PCR was positive in these patients.[8]
- The type of sample is an important factor. In patients with dry cough nasopharyngeal swabs should be preferred over oropharyngeal swabs.[9] The lower respiratory tract samples have high viral load and positive detection rates compared to upper respiratory tract samples. A study was done in China which compared the positive detection rates in different specimens from 205 symptomatic patients. The study showed a 93% positive rate in bronchoalveolar lavage fluids specimens, 72 % in sputum, 63 % in nasal, and 32 % in the pharyngeal swab.[10]
The comparison of chest CT scan and RT-PCR
- There are research studies that compared the accuracy, sensitivity, and specificity of chest CT scans with RT-PCR in diagnosing coronavirus illness.
- The RT-PCR test is the reference standard for the comparison.
- A retrospective study was performed by Ai et al. in 1049 suspected patients at the largest hospital in Wuhan, China. [11] Gietema et al. did a prospective study on 193 symptomatic patients in a hospital in Netherland.[12] The results from both these studies are tabulated below:
| Author | Total number of patients | Sensitivity % (95 % CI) | Specificity % (95% CI) | Positive predictive value % (95% CI) | Negative predictive value % (95% CI) |
|---|---|---|---|---|---|
| Ai et al | 1049 | 89.2(80.4-94.9) | 68.2(58.6-76.7) | 67.9(61.4-73.7) | 89.3(81.6-94) |
| Gietema et al | 193 | 97(95-98) | 25(22-30) | 65(62-68) | 83(76-89) |
- Both studies showed high sensitivity of chest CT scans in diagnosing coronavirus illness.
- The RT-PCR has a low positive rate. The sensitivity of chest CT scan is overestimated, and specificity is underestimated by using RT-PCR as a reference test.[11]
- In the early course of the illness, both the RT-PCR and Chest CT scans can have false-negative results.[13] Serial RT-PCR tests should be performed in patients with clinical symptoms suggestive of coronavirus illness.
- The chest CT scan can have positive findings in other respiratory illnesses including viral pneumonia.[12]
The Comparison of Serological Assays with RT-PCR
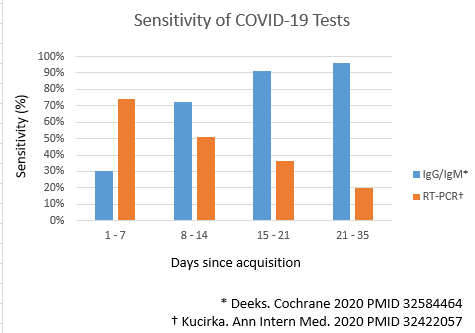
- The serological assays detect titers of antibodies against two structural proteins on SARS Cov-2. The proteins are S proteins from spike and N proteins from helical nucleocapsid of coronavirus.[14][15][16]Both these proteins are highly antigenic and elevated titers of antibodies are produced on exposure to it. IgM, IgG or both antibodies levels are measured by rapid lateral flow assays in serum, plasma, or whole blood samples.[17]
- The IgM and IgG antibodies rise in the later course of the illness.
- IgM antibodies titers rise first with a median duration of detection of 5 days after the onset of symptoms followed by a rise in IgG antibodies titer.
- The median time for the detection of IgG antibodies titers is 2 weeks after the onset of symptoms.[18]
- Sensitivity for IgG antibodies titers is over 90% after three weeks[19][20]
- The Serological assays can help detect subclinical, asymptomatic, late, and previous exposure to coronavirus. The serological assays increase the positive detection rates for detecting SARS Cov-2 when performed in conjunction with RT-PCR compared to PCR alone in suspected individuals. The positive detection rate of RT-PCR with IgM assay was 98.6% compared to 51.9% with PCR alone.[18]
- There are few advantages of serological assays over the PCR. The serological assays are less expensive, rapid, and easier to perform with less hazard of transmission to the virus to the health worker collecting the sample. [7]
- Secondly, the false-negative results due to sample characteristics in RT-PCR are absent in it. The antibodies are uniformly present in plasma. [18]
Diagnostic Criteria
The diagnostic criteria for suspected and confirmed cases of coronavirus disease 2019 (COVID-19) is tabulated below:[21][22][23][24][25][26]
| Case | Diagnostic Criteria |
|---|---|
| Suspected Case | Anyone with a history of epidemiology and any two of the clinical manifestations or anyone without epidemiological history and three of the clinical manifestations is considered to be a suspected case:
1) Epidemiological history:
|
| Confirmed Case | Any suspected case with one of the following pathogenic features is reclassified as a confirmed case:
|
Ongoing diagnostic Trials
- A researcher at Israel’s Ben-Gurion University of the Negev (BGU) has developed a test that identifies those carrying the COVID-19 virus in less than a minute. And it is both affordable and works with greater than 90% accuracy to boot[27]
Antibody response
- Most recovering from #COVID19 do not have high levels of neutralizing antibodies BUT antibodies to the receptor binding domain (RBD) of the spike protein with potent antiviral activity were found in all individuals tested & may be tx target[28]
- 8 weeks after hospital discharge, 40% of asymptomatic patients have no antibodies, and 12.9% of those who were symptomatic had no #COVID19 antibodies[29]
References
- ↑ Nalla AK, Casto AM, Huang MW, Perchetti GA, Sampoleo R, Shrestha L; et al. (2020). "Comparative Performance of SARS-CoV-2 Detection Assays Using Seven Different Primer-Probe Sets and One Assay Kit". J Clin Microbiol. 58 (6). doi:10.1128/JCM.00557-20. PMC 7269385 Check
|pmc=value (help). PMID 32269100 Check|pmid=value (help). - ↑ Tang YW, Schmitz JE, Persing DH, Stratton CW (2020). "Laboratory Diagnosis of COVID-19: Current Issues and Challenges". J Clin Microbiol. 58 (6). doi:10.1128/JCM.00512-20. PMC 7269383 Check
|pmc=value (help). PMID 32245835 Check|pmid=value (help). - ↑ Lieberman JA, Pepper G, Naccache SN, Huang ML, Jerome KR, Greninger AL (2020). "Comparison of Commercially Available and Laboratory Developed Assays for in vitro Detection of SARS-CoV-2 in Clinical Laboratories". J Clin Microbiol. doi:10.1128/JCM.00821-20. PMID 32350048 Check
|pmid=value (help). - ↑ Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H; et al. (2020). "First Case of 2019 Novel Coronavirus in the United States". N Engl J Med. 382 (10): 929–936. doi:10.1056/NEJMoa2001191. PMC 7092802 Check
|pmc=value (help). PMID 32004427 Check|pmid=value (help). - ↑ Corman VM, Landt O, Kaiser M, Molenkamp R, Meijer A, Chu DK; et al. (2020). "Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR". Euro Surveill. 25 (3). doi:10.2807/1560-7917.ES.2020.25.3.2000045. PMC 6988269 Check
|pmc=value (help). PMID 31992387. - ↑ Chan JF, Yip CC, To KK, Tang TH, Wong SC, Leung KH; et al. (2020). "Improved Molecular Diagnosis of COVID-19 by the Novel, Highly Sensitive and Specific COVID-19-RdRp/Hel Real-Time Reverse Transcription-PCR Assay Validated In Vitro and with Clinical Specimens". J Clin Microbiol. 58 (5). doi:10.1128/JCM.00310-20. PMC 7180250 Check
|pmc=value (help). PMID 32132196 Check|pmid=value (help). - ↑ 7.0 7.1 7.2 Zainol Rashid Z, Othman SN, Abdul Samat MN, Ali UK, Wong KK (2020). "Diagnostic performance of COVID-19 serology assays". Malays J Pathol. 42 (1): 13–21. PMID 32342927 Check
|pmid=value (help). - ↑ Fang, Yicheng; Zhang, Huangqi; Xie, Jicheng; Lin, Minjie; Ying, Lingjun; Pang, Peipei; Ji, Wenbin (2020). "Sensitivity of Chest CT for COVID-19: Comparison to RT-PCR". Radiology: 200432. doi:10.1148/radiol.2020200432. ISSN 0033-8419.
- ↑ Yu F, Yan L, Wang N, Yang S, Wang L, Tang Y; et al. (2020). "Quantitative Detection and Viral Load Analysis of SARS-CoV-2 in Infected Patients". Clin Infect Dis. doi:10.1093/cid/ciaa345. PMC 7184442 Check
|pmc=value (help). PMID 32221523 Check|pmid=value (help). - ↑ Wang W, Xu Y, Gao R, Lu R, Han K, Wu G; et al. (2020). "Detection of SARS-CoV-2 in Different Types of Clinical Specimens". JAMA. doi:10.1001/jama.2020.3786. PMC 7066521 Check
|pmc=value (help). PMID 32159775 Check|pmid=value (help). - ↑ 11.0 11.1 Ai T, Yang Z, Hou H, Zhan C, Chen C, Lv W; et al. (2020). "Correlation of Chest CT and RT-PCR Testing in Coronavirus Disease 2019 (COVID-19) in China: A Report of 1014 Cases". Radiology: 200642. doi:10.1148/radiol.2020200642. PMC 7233399 Check
|pmc=value (help). PMID 32101510 Check|pmid=value (help). - ↑ 12.0 12.1 Gietema HA, Zelis N, Nobel JM, Lambriks LJG, van Alphen LB, Oude Lashof AML; et al. (2020). "CT in relation to RT-PCR in diagnosing COVID-19 in The Netherlands: A prospective study". PLoS One. 15 (7): e0235844. doi:10.1371/journal.pone.0235844. PMID 32645053 Check
|pmid=value (help). - ↑ Xie X, Zhong Z, Zhao W, Zheng C, Wang F, Liu J (2020). "Chest CT for Typical 2019-nCoV Pneumonia: Relationship to Negative RT-PCR Testing". Radiology: 200343. doi:10.1148/radiol.2020200343. PMC 7233363 Check
|pmc=value (help). PMID 10.1148/radiol.2020200463 32049601 10.1148/radiol.2020200463 Check|pmid=value (help). - ↑ Chan CM, Tse H, Wong SS, Woo PC, Lau SK, Chen L; et al. (2009). "Examination of seroprevalence of coronavirus HKU1 infection with S protein-based ELISA and neutralization assay against viral spike pseudotyped virus". J Clin Virol. 45 (1): 54–60. doi:10.1016/j.jcv.2009.02.011. PMC 7108224 Check
|pmc=value (help). PMID 19342289. - ↑ Chan-Yeung M, Xu RH (2003). "SARS: epidemiology". Respirology. 8 Suppl: S9–14. doi:10.1046/j.1440-1843.2003.00518.x. PMC 7169193 Check
|pmc=value (help). PMID 15018127. - ↑ Liu Y, Eggo RM, Kucharski AJ (2020). "Secondary attack rate and superspreading events for SARS-CoV-2". Lancet. 395 (10227): e47. doi:10.1016/S0140-6736(20)30462-1. PMC 7158947 Check
|pmc=value (help). PMID 32113505 Check|pmid=value (help). - ↑ Li Z, Yi Y, Luo X, Xiong N, Liu Y, Li S; et al. (2020). "Development and clinical application of a rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis". J Med Virol. doi:10.1002/jmv.25727. PMC 7228300 Check
|pmc=value (help). PMID 32104917 Check|pmid=value (help). - ↑ 18.0 18.1 18.2 Guo L, Ren L, Yang S, Xiao M, Yang F; et al. (2020). "Profiling Early Humoral Response to Diagnose Novel Coronavirus Disease (COVID-19)". Clin Infect Dis. doi:10.1093/cid/ciaa310. PMC 7184472 Check
|pmc=value (help). PMID 32198501 Check|pmid=value (help). - ↑ Kucirka LM, Lauer SA, Laeyendecker O, Boon D, Lessler J (2020). "Variation in False-Negative Rate of Reverse Transcriptase Polymerase Chain Reaction-Based SARS-CoV-2 Tests by Time Since Exposure". Ann Intern Med. 173 (4): 262–267. doi:10.7326/M20-1495. PMC 7240870 Check
|pmc=value (help). PMID 32422057 Check|pmid=value (help). - ↑ Zhang JJY, Lee KS, Ong CW, Chan MY, Ang LW, Leo YS; et al. (2021). "Diagnostic performance of COVID-19 serological assays during early infection: A systematic review and meta-analysis of 11 516 samples". Influenza Other Respir Viruses. doi:10.1111/irv.12841. PMID 33609075 Check
|pmid=value (help). - ↑ Chen, Nanshan; Zhou, Min; Dong, Xuan; Qu, Jieming; Gong, Fengyun; Han, Yang; Qiu, Yang; Wang, Jingli; Liu, Ying; Wei, Yuan; Xia, Jia'an; Yu, Ting; Zhang, Xinxin; Zhang, Li (2020). "Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study". The Lancet. 395 (10223): 507–513. doi:10.1016/S0140-6736(20)30211-7. ISSN 0140-6736.
- ↑ "C.N.H. Commission Notice on prevention and control of pneumonia in children and pregnant women with new coronavirus infection China National Health Commission, Beijing (2020) (in Chinese)". line feed character in
|title=at position 18 (help) - ↑ "Technology, M.e.g.o.T.h.a.t.T.M.C.o.H.U.o.S.a. A rapid guideline for the diagnosis and treatment of pneumonia with new coronavirus infection (Third edition)".
- ↑ "Union Hospital T.M.C., Huazhong University of Science and Technology., Wuhan union hospital manage the 2019 new coronavirus infection strategies and instructions (in Chinese)". line feed character in
|title=at position 15 (help) - ↑ "[Diagnosis and clinical management of 2019 novel coronavirus infection: an operational recommendation of Peking Union Medical College Hospital (V2.0)]". Zhonghua Nei Ke Za Zhi (in Chinese). 59 (3): 186–188. February 2020. doi:10.3760/cma.j.issn.0578-1426.2020.03.003. PMID 32023681 Check
|pmid=value (help). - ↑ "Clinical management of severe acute respiratory infection when novel coronavirus (nCoV) infection is suspected".
- ↑ https://www.bioworld.com/articles/435285-Israels-ben-Gurion-university-develops-one-minute-coronavirus-test.html. Missing or empty
|title=(help) - ↑ Robbiani, Davide F.; Gaebler, Christian; Muecksch, Frauke; Lorenzi, Julio C. C.; Wang, Zijun; Cho, Alice; Agudelo, Marianna; Barnes, Christopher O.; Gazumyan, Anna; Finkin, Shlomo; Hägglöf, Thomas; Oliveira, Thiago Y.; Viant, Charlotte; Hurley, Arlene; Hoffmann, Hans-Heinrich; Millard, Katrina G.; Kost, Rhonda G.; Cipolla, Melissa; Gordon, Kristie; Bianchini, Filippo; Chen, Spencer T.; Ramos, Victor; Patel, Roshni; Dizon, Juan; Shimeliovich, Irina; Mendoza, Pilar; Hartweger, Harald; Nogueira, Lilian; Pack, Maggi; Horowitz, Jill; Schmidt, Fabian; Weisblum, Yiska; Michailidis, Eleftherios; Ashbrook, Alison W.; Waltari, Eric; Pak, John E.; Huey-Tubman, Kathryn E.; Koranda, Nicholas; Hoffman, Pauline R.; West, Anthony P.; Rice, Charles M.; Hatziioannou, Theodora; Bjorkman, Pamela J.; Bieniasz, Paul D.; Caskey, Marina; Nussenzweig, Michel C. (2020). "Convergent antibody responses to SARS-CoV-2 in convalescent individuals". Nature. doi:10.1038/s41586-020-2456-9. ISSN 0028-0836.
- ↑ Long, Quan-Xin; Tang, Xiao-Jun; Shi, Qiu-Lin; Li, Qin; Deng, Hai-Jun; Yuan, Jun; Hu, Jie-Li; Xu, Wei; Zhang, Yong; Lv, Fa-Jin; Su, Kun; Zhang, Fan; Gong, Jiang; Wu, Bo; Liu, Xia-Mao; Li, Jin-Jing; Qiu, Jing-Fu; Chen, Juan; Huang, Ai-Long (2020). "Clinical and immunological assessment of asymptomatic SARS-CoV-2 infections". Nature Medicine. doi:10.1038/s41591-020-0965-6. ISSN 1078-8956.