Nucleotide
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [1]
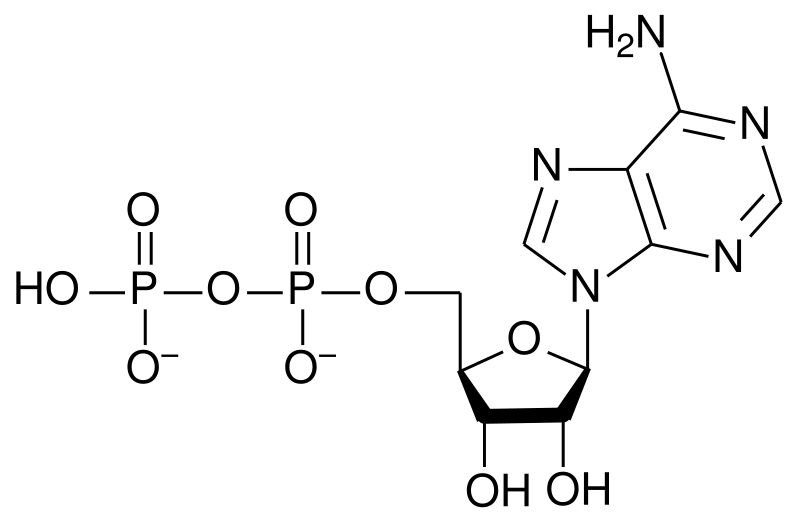
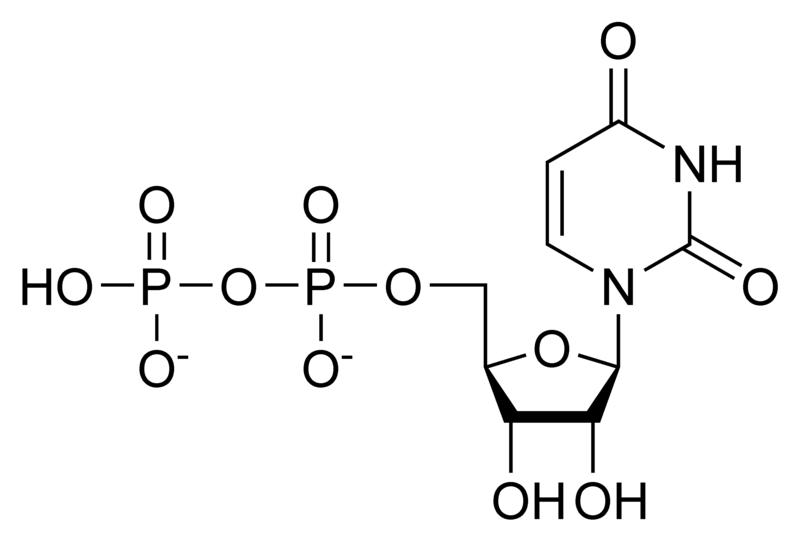
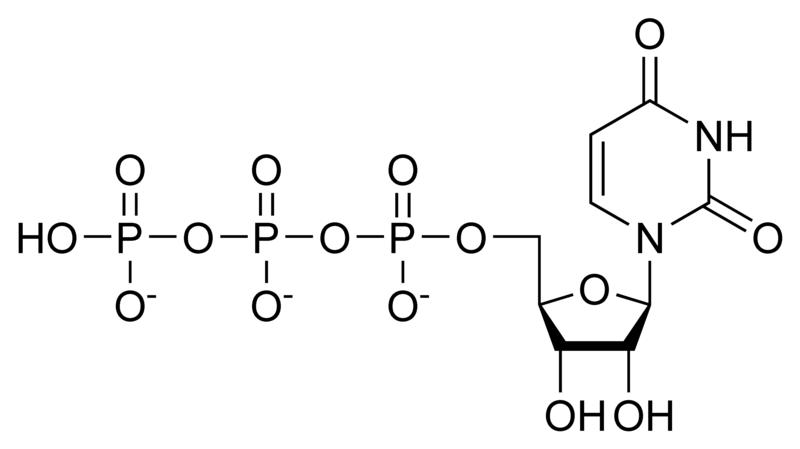
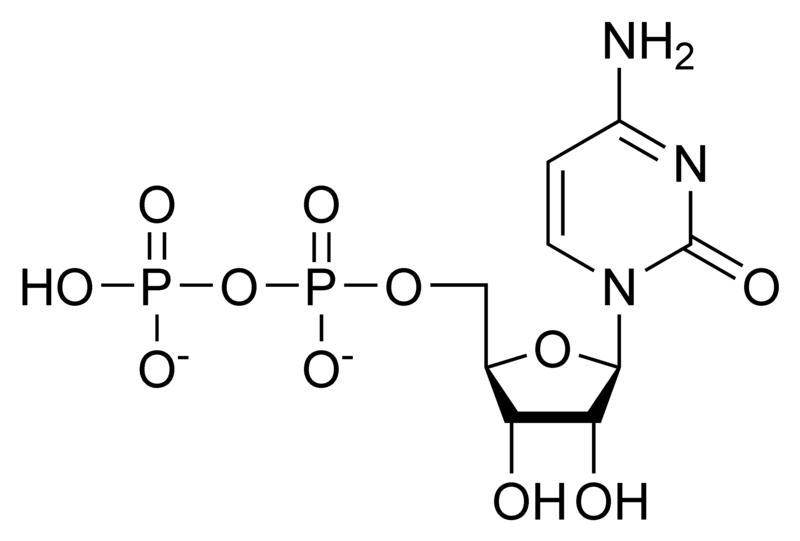
A nucleotide is a chemical compound that consists of 3 portions: a heterocyclic base, a sugar, and one or more phosphate groups. In the most common nucleotides the base is a derivative of purine or pyrimidine, and the sugar is the pentose (five-carbon sugar) deoxyribose or ribose. Nucleotides are the monomers of nucleic acids, with three or more bonding together in order to form a nucleic acid.
Nucleotides are the structural units of RNA, DNA, and several cofactors - CoA, flavin adenine dinucleotide, flavin mononucleotide, adenosine triphosphate and nicotinamide adenine dinucleotide phosphate. In the cell they have important roles in metabolism and signaling.

Nucleotides
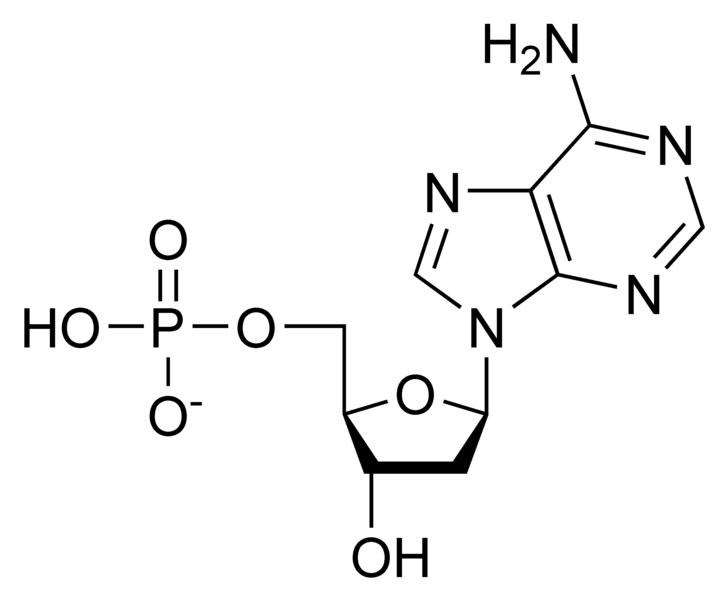
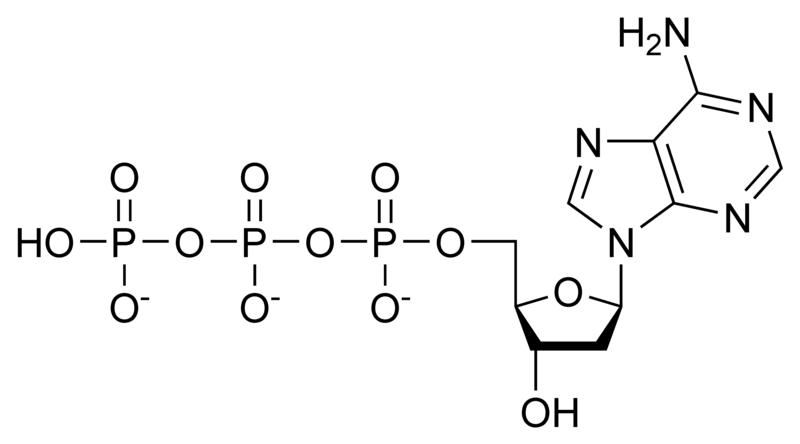
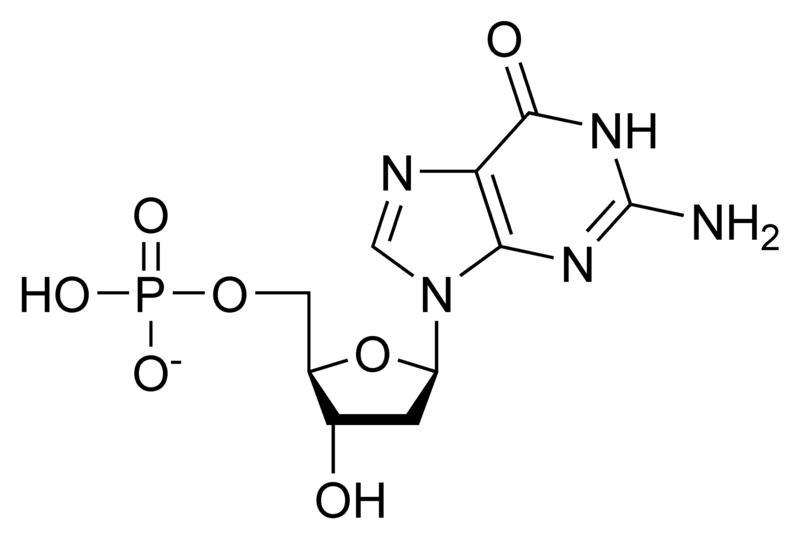
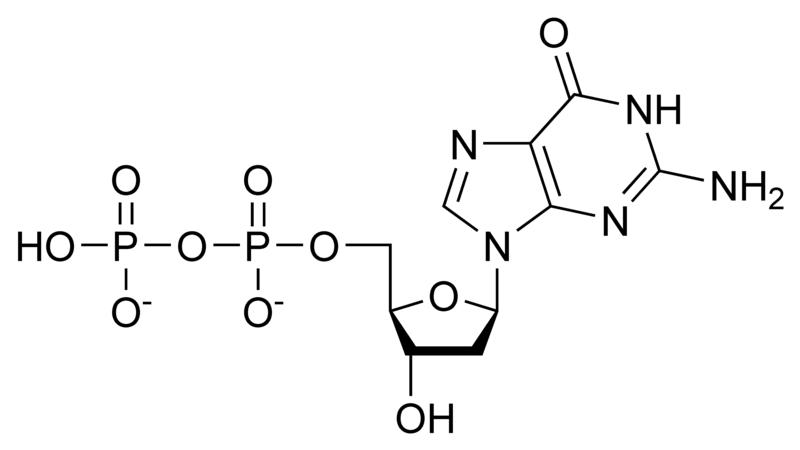
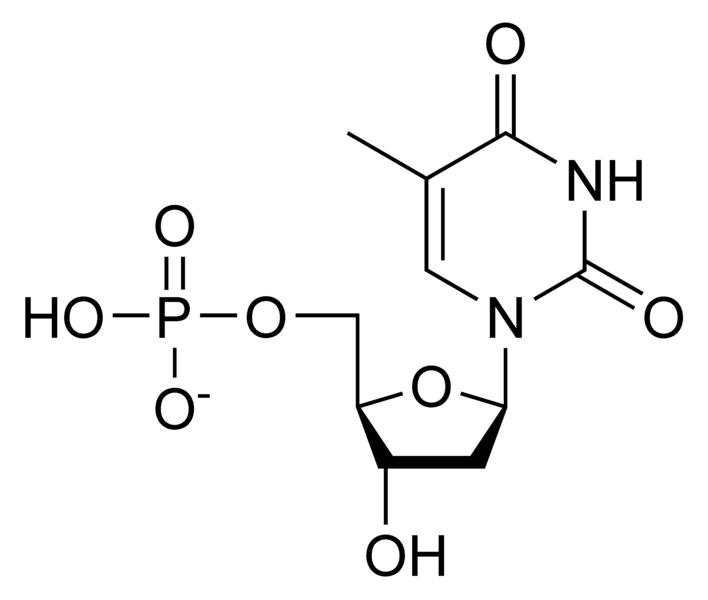
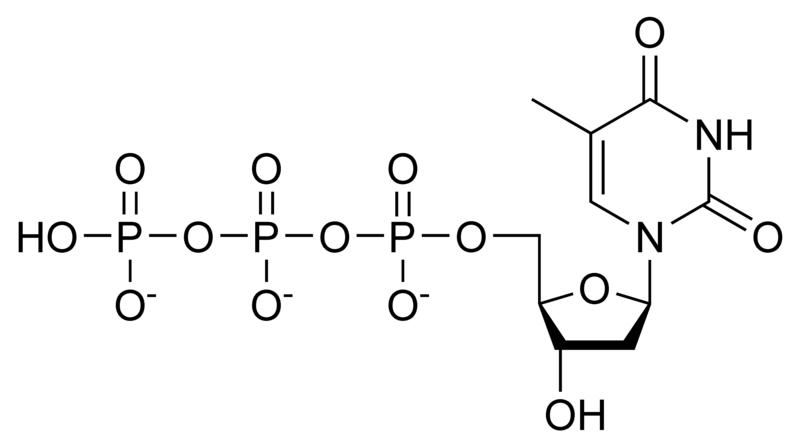
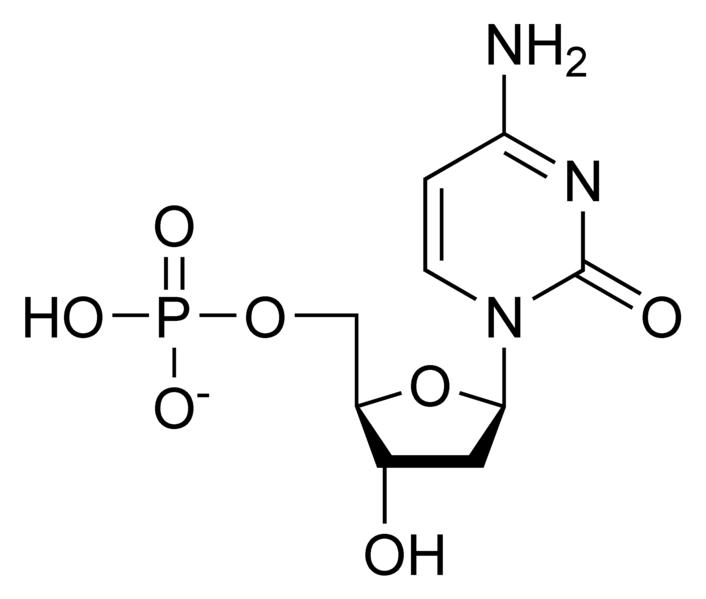
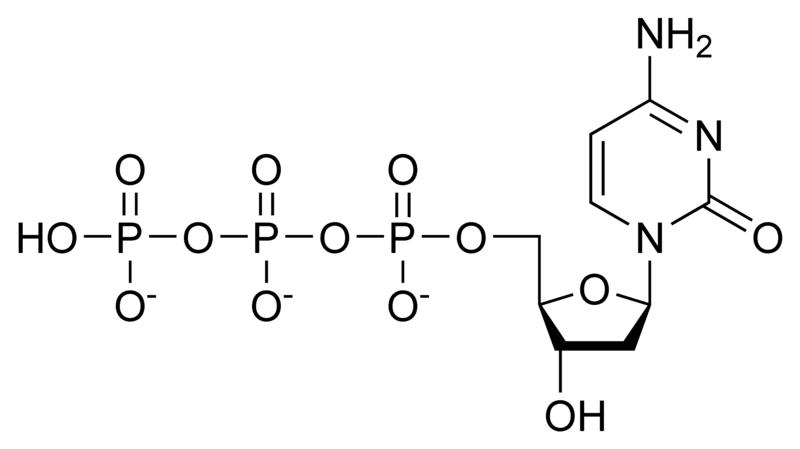
Deoxynucleotides
NOTE: If in place of ribose , the sugar deoxyribose is present the prefix `deoxy` may be added before the name of the nucleoside in all cases except thymidine.
Synthesis
Salvage synthesis refers to the reuse of parts of nucleotides in resynthesizing new nucleotides. Salvage synthesis requires both breakdown and synthesis reactions in order to exchange the useful parts.
Natural
Purine ribonucleotides
By using a variety of isotopically labelled compounds it was demonstrated that the sources of the atoms in purines are as follows:
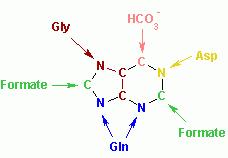 |
The biosynthetic origins of purine ring atoms N1 arises from the amine group of Asp C2 and C8 originate from formate N3 and N9 are contributed by the amide group of Gln C4, C5 and N7 are derived from Gly C6 comes from HCO3- (CO2) |

The de novo synthesis of purine nucleotides by which these precursors are incorporated into the purine ring, proceeds by a 10 step pathway to the branch point intermediate IMP, the nucleotide of the base hypoxanthine. AMP and GMP are subsequently synthesized from this intermediate via separate, two step each, pathways. Thus purine moieties are initially formed as part of the ribonucleotides rather than as free bases.
Six enzymes take part in IMP synthesis. Three of them are multifunctional:
Reaction 1. The pathway starts with the formation of PRPP. PRPS1 is the enzyme that activates R5P, which is primarily formed by the pentose phosphate pathway, to PRPP by reacting it with ATP. The reaction is unusual in that a pyrophosphoryl group is directly transferred from ATP to C1 of R5P and that the product has the α configuration about C1. This reaction is also shared with the pathways for the synthesis of the pyrimidine nucleotides, Trp, and His. As a result of being on (a) such (a) major metabolic crossroad and the use of energy, this reaction is highly regulated.
Reaction 2. In the first reaction unique to purine nucleotide biosynthesis, PPAT catalyzes the displacement of PRPP's pyrophosphate group (PPi) by Gln's amide nitrogen. The reaction occurs with the inversion of configuration about ribose C1, thereby forming β-5-phosphorybosylamine (5-PRA) and establishing the anomeric form of the future nucleotide. This reaction which is driven to completion by the subsequent hydrolysis of the released PPi, is the pathway's flux generating step and is therefore regulated too.
Reaction 3.
Pyrimidine ribonucleotides

Protection Chemistry
Nucleic acids can be synthesised in the lab. using protecting groups, typically this is achieved by protecting a purified nucleoside or nucleobase, a protected base is called a phosphoramidite. these can be used to obtain analogues not present in nature and/or to create an oligonucleotide.
References
See also
External links
- Abbreviations and Symbols for Nucleic Acids, Polynucleotides and their Constituents (IUPAC)
- Provisional Recommendations 2004 (IUPAC)
- Chemistry explanation of nucleotide structure
ar:نيوكليوتايد zh-min-nan:He̍k-kam-sng bs:Nukleotidi bg:Нуклеотид ca:Nucleòtid cs:Nukleotid da:Nukleotid de:Nukleotid el:Νουκλεοτίδιο eo:Nukleotido fa:نوکلئوتید gl:Nucleótido ko:뉴클레오티드 id:Nukleotida it:Nucleotide he:נוקלאוטיד ka:ნუკლეოტიდი lb:Nukleotid lt:Nukleotidas hu:Nukleotid ms:Nukleotida mk:Нуклеотид nl:Nucleotide simple:Nucleotide sk:Nukleotid sl:Nukleotid sr:Нуклеотид fi:Nukleotidi sv:Nukleotid th:นิวคลีโอไทด์ uk:Нуклеотиди ur:مرثالثہ