Antibiotic resistance
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [1]
Overview
Antibiotic resistance is the ability of a micro-organism to withstand the effects of an antibiotic. It is a specific type of drug resistance. Antibiotic resistance evolves naturally via natural selection through random mutation, but it could also be engineered. Once such a gene is generated, bacteria can then transfer the genetic information in a horizontal fashion (between individuals) by plasmid exchange. If a bacterium carries several resistance genes, it is called multiresistant or, informally, a superbug. Antibiotic resistance can also be introduced artificially into a micro-organism through transformation protocols. This can be a useful way of implanting artificial genes into the micro-organism.
Causes
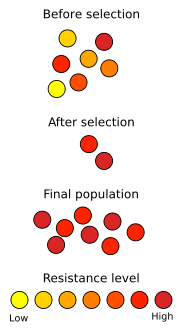
Antibiotic resistance is a consequence of evolution via natural selection. The antibiotic action is an environmental pressure; those bacteria which have a mutation allowing them to survive will live on to reproduce. They will then pass this trait to their offspring, which will be a fully resistant generation. If let untreated, the victim will quickly die.
Researchers have recently demonstrated the bacterial protein LexA may play a key role in the acquisition of bacterial mutations.
Studies have demonstrated that patterns of antibiotic usage greatly affect the number of resistant organisms which develop. Overuse of broad-spectrum antibiotics, such as second- and third-generation cephalosporins, greatly hastens the development of methicillin resistance. Other factors contributing towards resistance include incorrect diagnosis, unnecessary prescriptions, improper use of antibiotics by patients, and the use of antibiotics as livestock food additives for growth promotion.
Factors that affect how physicians prescribe antibiotics include:
- "Physicians were more likely to prescribe antibiotics to patients who they believed expected them, although they (the physicians) correctly identified only about 1 in 4 of those patients".
- Physicians may be more likely to prescribe antibiotics when the physicians are tired during clinical care.
- The use of antibiotics varies among countries.
Mechanisms
The four main mechanisms by which micro-organisms exhibit resistance to antimicrobials are:
- Drug inactivation or modification: e.g. enzymatic deactivation of Penicillin G in some penicillin-resistant bacteria through the production of β-lactamases.
- Alteration of target site : e.g. alteration of PBP—the binding target site of penicillins—in MRSA and other penicillin-resistant bacteria.
- Alteration of metabolic pathway: e.g. some sulfonamide-resistant bacteria do not require para-aminobenzoic acid (PABA), an important precursor for the synthesis of folic acid and nucleic acids in bacteria inhibited by sulfonamides. Instead, like mammalian cells, they turn to utilizing preformed folic acid.
- Reduced drug accumulation: by decreasing drug permeability and/or increasing active efflux on the cell surface.
Resistant pathogens
Staphylococcus aureus (colloquially known as "Staph aureus" or a Staph infection) is one of the major resistant pathogens. Found on the mucous membranes and the skin of around a third of the population, it is extremely adaptable to antibiotic pressure. It was the first bacterium in which penicillin resistance was found—in 1947, just four years after the drug started being mass-produced. Methicillin was then the antibiotic of choice, but has since been replaced by oxacillin due to significant kidney toxicity. MRSA (methicillin-resistant Staphylococcus aureus) was first detected in Britain in 1961 and is now "quite common" in hospitals. MRSA was responsible for 37% of fatal cases of blood poisoning in the UK in 1999, up from 4% in 1991. Half of all S. aureus infections in the US are resistant to penicillin, methicillin, tetracycline and erythromycin.
This left vancomycin as the only effective agent available at the time. However, VRSA (Vancomycin-resistant Staphylococcus aureus) was first identified in Japan in 1996, and has since been found in hospitals in England, France and the US. VRSA is also termed GISA (glycopeptide intermediate Staphylococcus aureus) or VISA (vancomycin insensitive Staphylococcus aureus), indicating resistance to all glycopeptide antibiotics.
A new class of antibiotics, oxazolidinones, became available in the 1990s, and the first commercially available oxazolidinone, linezolid, is comparable to vancomycin in effectiveness against MRSA. Linezolid-resistance in Staphylococcus aureus was reported in 2003.
CA-MRSA has now emerged as an epidemic that is responsible for rapidly progressive, fatal diseases including necrotizing pneumonia, severe sepsis and necrotizing fasciitis. Methicillin-resistant Staphylococcus aureus (MRSA) is the most frequently identified antimicrobial drug-resistant pathogen in US hospitals. The epidemiology of infections caused by MRSA is rapidly changing. In the past 10 years, infections caused by this organism have emerged in the community. The 2 MRSA clones in the United States most closely associated with community outbreaks, USA400 (MW2 strain, ST1 lineage) and USA300, often contain Panton-Valentine leukocidin (PVL) genes and, more frequently, have been associated with skin and soft tissue infections. Outbreaks of community-associated (CA)-MRSA infections have been reported in correctional facilities, among athletic teams, among military recruits, in newborn nurseries, and among active homosexual men. CA-MRSA infections now appear to be endemic in many urban regions and cause most CA-S. aureus infections.
Enterococcus faecium is another superbug found in hospitals. Penicillin-Resistant Enterococcus was seen in 1983, Vancomycin-Resistant Enterococcus (VRE) in 1987, and Linezolid-Resistant Enterococcus (LRE) in the late 1990s.
Streptococcus pyogenes (Group A Streptococcus: GAS) infections can usually be treated with many different antibiotics. Early treatment may reduce the risk of death from invasive group A streptococcal disease. However, even the best medical care does not prevent death in every case. For those with very severe illness, supportive care in an intensive care unit may be needed. For persons with necrotizing fasciitis, surgery often is needed to remove damaged tissue. Strains of S. pyogenes resistant to macrolide antibiotics have emerged, however all strains remain uniformly sensitive to penicillin.
Resistance of Streptococcus pneumoniae to penicillin and other beta-lactams is increasing worldwide. The major mechanism of resistance involves the introduction of mutations in genes encoding penicillin-binding proteins. Selective pressure is thought to play an important role, and use of beta-lactam antibiotics has been implicated as a risk factor for infection and colonization. Streptococcus pneumoniae is responsible for pneumonia, bacteremia, otitis media, meningitis, sinusitis, peritonitis and arthritis.
Proteus can cause urinary tract infections and hospital-acquired infections. Proteus is unique, however, because it is highly motile and does not form regular colonies. Instead, Proteus forms what are known as "swarming colonies" when plated on non-inhibitory media. The most important member of this genus is considered to be Proteus mirabilis, a cause of wound and urinary tract infections. Fortunately, most strains of Proteus mirabilis are sensitive to ampicillin and cephalosporins. Unlike its relative, Proteus vulgaris is not sensitive to these antibiotics. However, this organism is isolated less often in the laboratory and usually only targets immunosuppressed individuals. Proteus vulgaris occurs naturally in the intestines of humans and a wide variety of animals; also manure, soil and polluted waters. More than 80% of human urinary tract infections (UTI) are due to the bacterium Escherichia coli but urinary infections due to Proteus mirabilis are also well documented. Proteus mirabilis once attached to urinary tract, infects the kidney more commonly than E. coli. Proteus mirabilis belongs to family Enterobacteriaceae and is a gram-negative motile swarmer bacterium. Proteus mirabilis are often found as free-living organisms in soil and water but they are also parasitic in the upper urinary tract of human beings.
Penicillin-resistant pneumonia caused by Streptococcus pneumoniae (commonly known as pneumococcus), was first detected in 1967, as was penicillin-resistant gonorrhea. Resistance to penicillin substitutes is also known as beyond S. aureus. By 1993 Escherichia coli was resistant to five fluoroquinolone variants. Mycobacterium tuberculosis is commonly resistant to isoniazid and rifampin and sometimes universally resistant to the common treatments. Other pathogens showing some resistance include Salmonella, Campylobacter, and Streptococci.
In November 2004, the Centers for Disease Control and Prevention (CDC) reported an increasing number of Acinetobacter baumannii bloodstream infections in patients at military medical facilities in which service members injured in the Iraq/Kuwait region during Operation Iraqi Freedom and in Afghanistan during Operation Enduring Freedom were treated. Most of these showed multidrug resistance (MRAB), with a few isolates resistant to all drugs tested.
Pseudomonas
Pseudomonas aeruginosa is a highly relevant opportunistic pathogen. One of the most worrisome characteristics of P. aeruginosa consists in its low antibiotic susceptibility. This low susceptibility is attributable to a concerted action of multidrug efflux pumps with chromosomally-encoded antibiotic resistance genes and the low permeability of the bacterial cellular envelopes. Besides intrinsic resistance, P. aeruginosa easily develop acquired resistance either by mutation in chromosomally-encoded genes, either by the horizontal gene transfer of antibiotic resistance determinants. Development of multidrug resistance by P. aeruginosa isolates requires several different genetic events that include acquisition of different mutations and/or horizontal transfer of antibiotic resistance genes. Hypermutation favours the selection of mutation-driven antibiotic resistance in P. aeruginosa strains producing chronic infections, whereas the clustering of several different antibiotic resistance genes in integrons favours the concerted acquisition of antibiotic resistance determinants. Some recent studies have shown that phenotypic resistance associated to biofilm formation or to the emergence of small-colony-variants may be important in the response of P. aeruginosa populations to antibiotics treatment.
Role of animals
MRSA is acknowledged to be a human commensal and pathogen. MRSA has been found in cats, dogs and horses, where it can cause the same problems as it does in humans. Owners can transfer the organism to their pets and vice-versa, and MRSA in animals is generally believed to be derived from humans.
Currently, it is estimated that greater than 55% of the antibiotics used in the US are given to food animals (e.g. chickens, pigs and cattle) in the absence of disease. Antibiotic use in food animal production has been associated with the emergence of antibiotic-resistant strains of bacteria including Salmonella, Campylobacter, Escherichia coli and Enterococcus, among others. There is substantial evidence from the US and European Union that these resistant bacteria cause antibiotic-resistant infections in humans. The American Society for Microbiology (ASM), the American Public Health Association (APHA) and the American Medical Association (AMA) have called for substantial restrictions on antibiotic use in food animal production including an end to all non-therapeutic uses. The food animal and pharmaceutical industries have fought hard to prevent new regulations that would limit the use of antibiotics in food animal production. For example, in 2000 the US Food and Drug Administration (FDA) announced their intention to rescind approval for fluoroquinolone use in poultry production because of substantial evidence linking it to the emergence of fluoroquinolone resistant Campylobacter infections in humans. The final decision to ban fluoroquinolones from use in poultry production was not made until 5 years later because of challenges from the food animal and pharmaceutical industries. Today, there are two federal bills (S.742 and H.R. 2562) aimed at phasing out non-therapeutic antibiotics in US food animal production. These bills are endorsed by many public health and medical organizations including the American Nurses Association (ANA), the American Academy of Pediatrics (AAP), and the American Public Health Association (APHA).
The illegal use of amantadine to medicate poultry in the South of China and other parts of southeast Asia means that although the H5N1 (avian flu) strain that appeared in Hong Kong in 1997 was amantadine sensitive, the more recent strains have all been amantadine resistant. This seriously reduces the treatment options available to doctors in the event of an influenza pandemic.
Course duration and development of antibiotic resistance
- Completing the antibiotic course even after resolution of symptoms is widely believed to be a must to avoid the development of antibiotic resistant strains by bacteria. Dr.Martin J Llewelyn, the professor of infectious diseases at Brighton and Sussex Medical School investigated this concept further.
- The roots of this belief started back in the 1940s with the development of antibiotics. Alexander Fleming himself indicated in his Nobel prize speech that insufficient penicillin for treatment of streptococcal throat infection can cause other patients to get infected with resistant strains of the bacteria. But up till now, streptococci causing throat infection never developed resistance to penicillin after all these years, argues Dr.Llewelyn.
- Two mechanisms are responsible for antibiotic resistance: Targeted selection and collateral selection.
- An example of targeted selection is the resistance developed by Mycobacterium tuberculosis when an insufficient or a single antibiotic is administered. This accounts for a minority of cases of antibiotic resistance.[1]
- Collateral selection occurs when the bacterial flora present normally in the body is affected by the administered antibiotic leaving space for other resistant harmful bacterial strains.
- Collateral selection accounts for most of the cases of antibiotic resistance and it is not dependent at all on the duration of administration of the antibiotic.
- The role of the duration of antibiotic administration on antibiotic resistance is clearly understudied and for most bacterial infections. The conventional course duration was not compared to a shorter course for efficacy and the development of resistant strains in most of the indications of administration of antibiotics.
- Pyelonephritis was treated with quinolones for two weeks, however, trials have shown that 5-7 day course is as efficient.
- Antibiotic treatment efficacy for otitis media was not the same with shorter courses. Antibiotics gave the same clinical results with shorter and longer treatment courses and in fact, shorter courses were associated with lower recurrence rates.
- As regarding hospital acquired pneumonia, research has shown that shorter antibiotic courses might be associated with less chance of developing resistant bacterial strains in the future.
- None of these trials have shown that antibiotic resistance was associated with the shorter courses.
- The author suggested administering antibiotics should be guided by infection biomarkers such as procalcitonin in the inpatient setting and with the symptoms of the patient in the outpatient setting. He also suggests that the message of “completing the course” should be reconsidered and changed.
- However, the topic needs further study to confirm this theory.
Alternatives
Prevention
Washing hands properly reduces the chance of getting infected or spreading infection. Thoroughly washing or avoiding raw foods such as fruits, vegetables, raw eggs, and undercooked meat can also reduce the chance of an infection. High risk activities include: unprotected sex ; use of equipment in a public gymnasium or on a playground; being a patient in a nursing home or hospital; being a prison inmate; visiting a barbershop; the sharing of personal products (cosmetic items, lotion, bedding, toothpaste, headphones, nail clippers, shampoo); even being overweight.
Avoiding the use of antibiotics, in some situations, can also reduce the risk of infection by antibiotic-resistant bacteria. One study demonstrated that the use of fluoroquinolones is clearly associated with elevated rates of Clostridium difficile infections, which is a leading cause of nosocomial diarrhea in the United States and a major cause of death worldwide.
Interventions to reduce over-prescription of antibiotics have been systematically reviewed. Multifactorial interventions aimed at both physicians and patients can reduce inappropriate prescription of antibiotics. The delay of antibiotics administration for 48 hours while waiting on improvement of respiratory tract infections or cystitis may reduce antibiotic usage and re-consultation; however, this strategy may reduce patient satisfaction.
The rate of prescription of antibiotics may be reduced with the reassurance of clinicians that serious infection is not present. Health care providers can be reassured when there is any of the following:
- Normal c-reactive protein levels
- Normal procalcitonin levels
Vaccines
Vaccines do not suffer the problem of resistance because a vaccine enhances the body's natural defenses, while an antibiotic operates separately from the body's normal defenses. Nevertheless, new strains may evolve that escape immunity induced by vaccines.
While theoretically promising, anti-staphylococcal vaccines have shown limited efficacy, because of immunological variation between Staphylococcus species, and the limited duration of effectiveness of the antibodies produced. Development and testing of more effective vaccines is under way.
Phage therapy
Phage therapy, an approach that has been extensively researched and utilized as a therapeutic agent for over 60 years, especially in the Soviet Union, is an alternative that might help with the problem of resistance. Phage Therapy was widely used in the United States until the discovery of antibiotics, in the early 1940's. Bacteriophages or "phages" are viruses that invade bacterial cells and, in the case of lytic phages, disrupt bacterial metabolism and cause the bacterium to lyse [destruct]. Phage Therapy is the therapeutic use of lytic bacteriophages to treat pathogenic bacterial infections.
Bacteriophage therapy is an important alternative to antibiotics in the current era of multidrug resistant pathogens. A review of studies that dealt with the therapeutic use of phages from 1966–1996 and few latest ongoing phage therapy projects via internet showed: phages were used topically, orally or systemically in Polish and Soviet studies. The success rate found in these studies was 80–95% with few gastrointeslinal or allergic side effects. British studies also demonstrated significant efficacy of phages against Escherichia coli, Acinetobacter spp., Pseudomonas spp and Staphylococcus aureus. US studies dealt with improving the bioavailability of phage. Phage therapy may prove as an important alternative to antibiotics for treating multidrug resistant pathogens.
Development of new drugs
Until recently, research and development (R&D) efforts have provided new drugs in time to treat bacteria that became resistant to older antibiotics. That is no longer the case. The potential crisis at hand is the result of a marked decrease in industry R&D, government inaction, and the increasing prevalence of resistant bacteria. Infectious disease physicians are alarmed by the prospect that effective antibiotics may not be available to treat seriously ill patients in the near future.
The pipeline of new antibiotics is drying up. Major pharmaceutical companies are losing interest in the antibiotics market because these drugs may not be as profitable as drugs that treat chronic (long-term) conditions and lifestyle issues.
The resistance problem demands that a renewed effort be made to seek antibacterial agents effective against pathogenic bacteria resistant to current antibiotics. One of the possible strategies towards this objective is the rational localization of bioactive phytochemicals. Plants have an almost limitless ability to synthesize aromatic substances, most of which are phenols or their oxygen-substituted derivatives such as tannins. Most are secondary metabolites, of which at least 12,000 have been isolated, a number estimated to be less than 10% of the total. In many cases, these substances serve as plant defense mechanisms against predation by micro-organisms, insects, and herbivores. Many of the herbs and spices used by humans to season food yield useful medicinal compounds including those having antibacterial activity
Traditional healers have long used plants to prevent or cure infectious conditions. Many of these plants have been investigated scientifically for antimicrobial activity and a large number of plant products have been shown to inhibit growth of pathogenic bacteria. A number of these agents appear to have structures and modes of action that are distinct from those of the antibiotics in current use, suggesting that cross-resistance with agents already in use may be minimal. For example the combination of 5'-methoxyhydnocarpine and berberine in herbs like Hydrastis canadensis and Berberis vulgaris can block the MDR-pumps that cause multidrug resistance. This has been shown for Staphylococcus aureus..
Applications
Antibiotic resistance is an important tool for genetic engineering. By constructing a plasmid which contains an antibiotic resistance gene as well as the gene being engineered or expressed, a researcher can ensure that when bacteria replicate, only the copies which carry along the plasmid survive. This ensures that the gene being manipulated passes along when the bacteria replicates.
The most commonly used antibiotics in genetic engineering are generally "older" antibiotics which have largely fallen out of use in clinical practice. These include:
Industrially the use of antibiotic resistance is disfavored since maintaining bacterial cultures would require feeding them large quantities of antibiotics. Instead, the use of auxotrophic bacterial strains (and function-replacement plasmids) is preferred.
See also
- LexA
- Efflux
- List of environment topics
- Nosocomial infection
- Tuberculosis
- Bacterial conjugation
- Drug of last resort
External links
- MRSA Action UK Registered Charity No. 1115672 - Supporting sufferers, dependants and carers in the UK
- CDC Article on Hospital Acquired MRSA
- CDC Article on Community Acquired MRSA
- CDC Guideline "Management of Multidrug-Resistant Organisms in Healthcare Settings, 2006"
- Vancomycin Resistant Enterococcus—Guidelines for Healthcare Workers
- Alliance for the Prudent Use of Antibiotics
- Quantifying potential human health impacts of animal antibiotic use: Enrofloxacin and macrolides in chickens.
- Rise of Antibiotic-Resistant Infections
- Information about phage therapy – a possible alternative to antibiotics in case of resistant infections
- Antibiotic-resistance genes as markers Once necessary, now undesirable
- CBS Article on Phage Therapy and Antibiotic Resistance
- BURDEN of Resistance and Disease in European Nations - An EU-Project to estimate the financial burden of antibiotic resistance in European Hospitals
References
- Lord Soulsby of Swaffham Prior (2005). "Resistance to antimicrobials in humans and animals". Brit J Med. 331: 1219&ndash, 20.
da:Antibiotikaresistens de:Antibiotikum-Resistenz nl:Resistentie tegen antibiotica sv:Antibiotikaresistens uk:Антибіотичний опір