Adenosine triphosphate
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [3]
Overview
Adenosine 5'-triphosphate (ATP) is a multifunctional nucleotide that is most important as a "molecular currency" of intracellular energy transfer.[1] In this role, ATP transports chemical energy within cells for metabolism. It is produced as an energy source during the processes of photosynthesis and cellular respiration and consumed by many enzymes and a multitude of cellular processes including biosynthetic reactions, motility and cell division. In signal transduction pathways, ATP is used as a substrate by kinases that phosphorylate proteins and lipids, as well as by adenylate cyclase, which uses ATP to produce the second messenger molecule cyclic AMP.
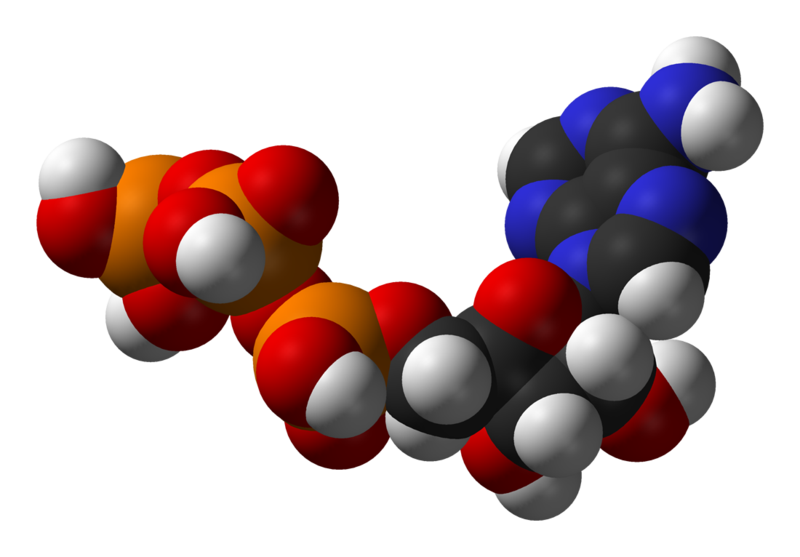
The structure of this molecule consists of a purine base (adenine) attached to the 1' carbon atom of a pentose sugar (ribose). Three phosphate groups are attached at the 5' carbon atom of the pentose sugar. ATP is also incorporated into nucleic acids by polymerases in the processes of DNA replication and transcription. When ATP is used in DNA synthesis, the ribose sugar is first converted to deoxyribose by ribonucleotide reductase. ATP was discovered in 1929 by Karl Lohmann,[2] and was proposed to be the main energy-transfer molecule in the cell by Fritz Albert Lipmann in 1941.[3]
Physical and chemical properties
ATP consists of adenosine — itself composed of an adenine ring and a ribose sugar — and three phosphate groups (triphosphate). The phosphoryl groups, starting with the group closest to the ribose, are referred to as the alpha (α), beta (β), and gamma (γ) phosphates. ATP is highly soluble in water and is quite stable in solutions between pH 6.8–7.4, but is rapidly hydrolysed at extreme pH, consequently ATP is best stored as an anhydrous salt.[4]
As ATP is an unstable molecule it tends to be hydrolysed in water, and if ATP and ADP are allowed to come to chemical equilibrium, almost all the ATP will be converted to ADP. Any system that is far from equilibrium contains potential energy, and is capable of doing work. The cell maintains the ratio of ATP to ADP at a point ten orders of magnitude from equilibrium, with ATP concentrations a thousandfold higher than the concentration of ADP. This displacement from equilibrium means that the hydrolysis of ATP in the cell releases a great deal of energy.[5] ATP is commonly referred to as a "high energy molecule", however this is incorrect, as a mixture of ATP and ADP at equilibrium in water can do no useful work at all. In fact, ATP does not contain any special "high-energy bonds" and any other unstable molecule would serve equally well as a way of storing energy if the cell maintained its concentration far from equilibrium.
The amount of energy released can be calculated from the changes in energy under non-natural conditions. The net change in heat energy (enthalpy) at standard temperature and pressure of the decomposition of ATP into hydrated ADP and hydrated inorganic phosphate is −20.5 kJ/mol, with a change in free energy of 3.4 kJ/mol.[6] The energy released by cleaving either a phosphate (Pi) or pyrophosphate (PPi) unit from ATP, with all reactants and products at their standard states of 1 M concentration, are:
- ATP + H2O → ADP(hydrated) + Pi(hydrated) + H+(hydrated) ΔG˚ = -30.54 kJ/mol (−7.3 kcal/mol)
- ATP + H2O → AMP(hydrated) + PPi(hydrated) + H+(hydrated) ΔG˚ = -45.6 kJ/mol (−10.9 kcal/mol)
These values can be used to calculate the change in energy under physiological conditions and the cellular ATP/ADP ratio. Note the values given for the Gibbs free energy for this reaction are dependent on a number of factors, including overall ionic strength and the presence of alkaline earth metal ions such as Mg2+ and Ca2+. Under typical cellular conditions, ΔG is approximately −57 kJ/mol (−14 kcal/mol).[7]
Ionization in biological systems
ATP has multiple ionizable groups with different acid dissociation constants. In neutral solution, ATP is ionized and exists mostly as ATP4−, with a small proportion of ATP3−.[8] As ATP has several negatively-charged groups in neutral solution, it can chelate metals with very high affinity. The binding constant for various metal ions are (given as per mole) as Mg2+ (9 554), Na+ (13), Ca2+ (3 722), K+ (8), Sr2+ (1 381) and Li+ (25).[9] Due to the strength of these interactions, ATP exists in the cell mostly in a complex with Mg2+.[10][8]
Biosynthesis
The ATP concentration inside the cell is typically 1 - 10 mM.[11] ATP can be produced by redox reactions using simple and complex sugars (carbohydrates) or lipids as an energy source. For ATP to be synthesized from complex fuels, they first need to be broken down into their basic components. Carbohydrates are hydrolysed into simple sugars, such as glucose and fructose. Fats (triglycerides) are metabolised to give fatty acids and glycerol.
The overall process of oxidizing glucose to carbon dioxide is known as cellular respiration and can produce up to 36 molecules of ATP from a single molecule of glucose.[12] ATP can be produced by a number of distinct cellular processes; the three main pathways used to generate energy in eukaryotic organisms are glycolysis and the citric acid cycle/oxidative phosphorylation , both components of cellular respiration; and beta-oxidation. The majority of this ATP production by a non-photosynthetic aerobic eukaryote takes place in the mitochondria, which can make up nearly 25% of the total volume of a typical cell.[12]
Glycolysis
In glycolysis, glucose and glycerol are metabolized to pyruvate via the glycolytic pathway. In most organisms this process occurs in the cytosol, but in some protozoa such as the kinetoplastids, this is carried out in a specialized organelle called the glycosome.[13] Glycolysis generates a net two molecules of ATP through substrate phosphorylation catalyzed by two enzymes: PGK and pyruvate kinase. Two molecules of NADH are also produced, which can be oxidized via the electron transport chain and result in the generation of additional ATP by ATP synthase. The pyruvate generated as an end-product of glycolysis is a substrate for the Krebs Cycle.
Citric acid cycle
In the mitochondrion, pyruvate is oxidized by the pyruvate dehydrogenase complex to acetyl CoA, which is fully oxidized to carbon dioxide by the citric acid cycle (also known as the Krebs Cycle). Every "turn" of the citric acid cycle produces two molecules of carbon dioxide, one molecule of the ATP equivalent guanosine triphosphate (GTP) through substrate-level phosphorylation catalyzed by succinyl CoA synthetase, three molecules of the reduced coenzyme NADH, and one molecule of the reduced coenzyme FADH2. Both of these latter molecules are recycled to their oxidized states (NAD+ and FAD, respectively) via the electron transport chain, which generates additional ATP by oxidative phosphorylation. The oxidation of an NADH molecule results in the synthesis of about 3 ATP molecules, and the oxidation of one FADH2 yields about 2 ATP molecules.[14] The majority of cellular ATP is generated by this process. Although the citric acid cycle itself does not involve molecular oxygen, it is an obligately aerobic process because O2 is needed to recycle the reduced NADH and FADH2 to their oxidized states. In the absence of oxygen the citric acid cycle will cease to function due to the lack of available NAD+ and FAD.[12]
The generation of ATP by the mitochondrion from cytosolic NADH relies on the malate-aspartate shuttle (and to a lesser extent, the glycerol-phosphate shuttle) because the inner mitochondrial membrane is impermeable to NADH and NAD+. Instead of transferring the generated NADH, a malate dehydrogenase enzyme converts oxaloacetate to malate, which is translocated to the mitochondrial matrix. Another malate dehydrogenase-catalyzed reaction occurs in the opposite direction, producing oxaloacetate and NADH from the newly transported malate and the mitochondrion's interior store of NAD+. A transaminase converts the oxaloacetate to aspartate for transport back across the membrane and into the intermembrane space.[12]
In oxidative phosphorylation, the passage of electrons from NADH and FADH2 through the electron transport chain powers the pumping of protons out of the mitrochondrial matrix and into the intermembrane space. This creates a proton motive force that is the net effect of a pH gradient and an electric potential gradient across the inner mitochondrial membrane. Flow of protons down this potential gradient — that is, from the intermembrane space to the matrix — provides the driving force for ATP synthesis by ATP synthase. This enzyme contains a rotor subunit that physically rotates relative to the static portions of the protein during ATP synthesis.[15]
Most of the ATP synthesized in the mitochondria will be used for cellular processes in the cytosol; thus it must be exported from its site of synthesis in the mitochondrial matrix. The inner membrane contains an antiporter, the ADP/ATP translocase, which is an integral membrane protein used to exchange newly-synthesized ATP in the matrix for ADP in the intermembrane space.[16] This translocase is driven by the membrane potential, as it results in the movement of about 4 negative charges out of the mitochondrial membrane in exchange for 3 negative charges moved inside. However, it is also necessary to transport phosphate into the mitochondrion; the phosphate carrier moves a proton in with each phosphate, partially dissipating the proton gradient.
Beta-oxidation
Fatty acids can also be broken down to acetyl-CoA by beta-oxidation. Each turn of this cycle reduces the length of the acyl chain by two carbon atoms and produces one NADH and one FADH2 molecule, which are used to generate ATP by oxidative phosphorylation. Because NADH and FADH2 are energy-rich molecules, dozens of ATP molecules can be generated by the beta-oxidation of a single long acyl chain. The high energy yield of this process and the compact storage of fat explain why it is the most dense source of dietary calories.[17]
Anaerobic respiration
Anaerobic respiration or fermentation entails the generation of energy via the process of oxidation in the absence of O2 as an electron acceptor. In most eukaryotes, glucose is used as both an energy store and an electron donor. The equation for the oxidation of glucose to lactic acid is:
- C6H12O6<math>\to</math> 2CH3CH(OH)COOH + 2 ATP
In prokaryotes, multiple electron acceptors can be used in anaerobic respiration. These include nitrate, sulfate or carbon dioxide. These processes lead to the ecologically-important processes of denitrification, sulfate reduction and acetogenesis, respectively.[18][19]
ATP replenishment by nucleoside diphosphate kinases
ATP can also be synthesized through several so-called "replenishment" reactions catalyzed by the enzyme families of nucleoside diphosphate kinases (NDKs), which use other nucleoside triphosphates as a high-energy phosphate donor, and the ATP:guanido-phosphotransferase family, which uses creatine.
ATP production during photosynthesis
In plants, ATP is synthesized in thylakoid membrane of the chloroplast during the light-dependent reactions of photosynthesis in a process called photophosphorylation. Here, light energy is used to pump protons across the chloroplast membrane. This produces a proton-motive force and this drives the ATP synthase, exactly as in oxidative phosphorylation.[20] Some of the ATP produced in the chloroplasts is consumed in the Calvin cycle, which produces triose sugars.
ATP recycling
The total quantity of ATP in the human body is about 0.1 mole. The majority of ATP is not usually synthesised de novo, but is generated from ADP by the aforementioned processes. Thus, at any given time, the total amount of ATP + ADP remains fairly constant.
The energy used by human cells requires the hydrolysis of 100 to 150 moles of ATP daily which is around 50 to 75 kg. Typically, a human will use up their body weight of ATP over the course of the day.[21] This means that each ATP molecule is recycled 1000 to 1500 times during a single day (100 / 0.1 = 1000). ATP cannot be stored, hence its consumption closely follows its synthesis.
Regulation of biosynthesis
ATP production in an aerobic eukaryotic cell is tightly regulated by allosteric mechanisms, by feedback effects, and by the substrate concentration dependence of individual enzymes within the glycolysis and oxidative phosphorylation pathways. Key control points occur in enzymatic reactions that are so energetically favorable that they are effectively irreversible under physiological conditions.
In glycolysis, hexokinase is directly inhibited by its product, glucose-6-phosphate, and pyruvate kinase is inhibited by ATP itself. The main control point for the glycolytic pathway is phosphofructokinase (PFK), which is allosterically inhibited by high concentrations of ATP and activated by high concentrations of AMP. The inhibition of PFK by ATP is unusual, since ATP is also a substrate in the reaction catalyzed by PFK; the biologically active form of the enzyme is a tetramer that exists in two possible conformations, only one of which binds the second substrate fructose-6-phosphate (F6P). The protein has two binding sites for ATP - the active site is accessible in either protein conformation, but ATP binding to the inhibitor site stabilizes the conformation that binds F6P poorly.[14] A number of other small molecules can compensate for the ATP-induced shift in equilibrium conformation and reactivate PFK, including cyclic AMP, ammonium ions, inorganic phosphate, and fructose 1,6 and 2,6 biphosphate.[14]
The citric acid cycle is regulated mainly by the availability of key substrates, particularly the ratio of NAD+ to NADH and the concentrations of calcium, inorganic phosphate, ATP, ADP, and AMP. Citrate - the molecule that gives its name to the cycle - is a feedback inhibitor of citrate synthase and also inhibits PFK, providing a direct link between the regulation of the citric acid cycle and glycolysis.[14]
In oxidative phosphorylation, the key control point is the reaction catalyzed by cytochrome c oxidase, which is regulated by the availability of its substrate—the reduced form of cytochrome c. The amount of reduced cytochrome c available is directly related to the amounts of other substrates:
- <math>
\frac{1}{2}NADH + cyt~c_{ox} + ADP + P_{i} \iff \frac{1}{2}NAD^{+} + cyt~c_{red} + ATP </math> which directly implies this equation:
- <math>
\frac{cyt~c_{red}}{cyt~c_{ox}} = \left(\frac{[NADH]}{[NAD]^{+}}\right)^{\frac{1}{2}}\left(\frac{[ADP][P_{i}]}{[ATP]}\right)K_{eq} </math>
Thus, a high ratio of [NADH] to [NAD+] or a low ratio of [ADP][Pi] to [ATP] imply a high amount of reduced cytochrome c and a high level of cytochrome c oxidase activity.[14] An additional level of regulation is introduced by the transport rates of ATP and NADH between the mitochondrial matrix and the cytoplasm.[16]
Functions in cells
ATP is generated in the cell by energy-releasing processes and is broken down by energy-consuming processes, in this way ATP transfers energy between spatially-separate metabolic reactions. ATP is the main energy source for the majority of cellular functions. This includes the synthesis of macromolecules, including DNA, RNA, and proteins. ATP also plays a critical role in the transport of macromolecules across cell membranes, e.g. exocytosis and endocytosis.
In the synthesis of the nucleic acid RNA, ATP is one of the four nucleotides incorporated directly into RNA molecules by RNA polymerases. The energy driving this polymerization comes from cleaving off a pyrophosphate (two phosphate groups).[22] The process is similar in DNA biosynthesis, except that ATP is reduced to the deoxyribonucleotide dATP, before incorporation into DNA.
ATP is critically involved in maintaining cell structure by facilitating assembly and disassembly of elements of the cytoskeleton. In a related process, ATP is required for the shortening of actin and myosin filament crossbridges required for muscle contraction. This latter process is one of the main energy requirements of animals and is essential for locomotion and respiration.
Cell signaling
Extracellular signaling
ATP is also a signaling molecule. ATP, ADP, or adenosine are recognized by purinergic receptors.
In humans, this signaling role is important in both the central and peripheral nervous system. Activity-dependent release of ATP from synapses, axons and glia activates purinergic membrane receptors known as P2.[23] The P2Y receptors are metabotropic, i.e. G protein-coupled and modulate mainly intracellular calcium and sometimes cyclic AMP levels. Fifteen members of the P2Y family have been reported (P2Y1–P2Y15), although some are only related through weak homology and several (P2Y5, P2Y7, P2Y9, P2Y10) do not function as receptors that raise cytosolic calcium. The P2X ionotropic receptor subgroup comprises seven members (P2X1–P2X7) which are ligand-gated Ca2+-permeable ion channels that open when bound to an extracellular purine nucleotide. In contrast to P2 receptors (agonist order ATP > ADP > AMP > ADO), purinergic nucleotides like ATP are not strong agonists of P1 receptors which are strongly activated by adenosine and other nucleosides (ADO > AMP > ADP > ATP). P1 receptors have A1, A2a, A2b, and A3 subtypes ("A" as a remnant of old nomenclature of adenosine receptor), all of which are G protein-coupled receptors, A1 and A3 being coupled to Gi, and A2a and A2b being coupled to Gs.[24]
Intracellular signaling
ATP is critical in signal transduction processes. It is used by kinases as the source of phosphate groups in their phosphate transfer reactions. Kinase activity on substrates such as proteins or membrane lipids are a common form of signal transduction. Phosphorylation of a protein by a kinase can activate this cascade such as the mitogen-activated protein kinase cascade.[25]
ATP is also used by adenylate cyclase and is transformed to the second messenger molecule cyclic AMP, which is involved in triggering calcium signals by the release of calcium from intracellular stores.[26] This form of signal transduction is particularly important in brain function, although it is involved in the regulation of a multitude of other cellular processes.[27]
Deoxyribonucleotide synthesis
In all known organisms, the deoxyribonucleotides that make up DNA are synthesized by the action of ribonucleotide reductase (RNR) enzymes on their corresponding ribonucleotides.[28] This enzyme reduces the 2' hydroxyl group on the ribose sugar to deoxyribose, forming a deoxyribonucleotide (denoted dATP). All ribonucleotide reductase enzymes use a common sulfhydryl radical mechanism reliant on reactive cysteine residues that oxidize to form disulfide bonds in the course of the reaction.[28] RNR enzymes are recycled by reaction with thioredoxin or glutaredoxin.[14]
The regulation of RNR and related enzymes maintains a balance of dNTPs relative to each other and relative to NTPs in the cell. Very low dNTP concentration inhibits DNA synthesis and DNA repair and is lethal to the cell, while an abnormal ratio of dNTPs is mutagenic due to the increased likelihood of the DNA polymerase incorporating the wrong dNTP during DNA synthesis.[14] Regulation of or differential specificity of RNR has been proposed as a mechanism for alterations in the relative sizes of intracellular dNTP pools under cellular stress such as hypoxia.[29]
Binding to proteins
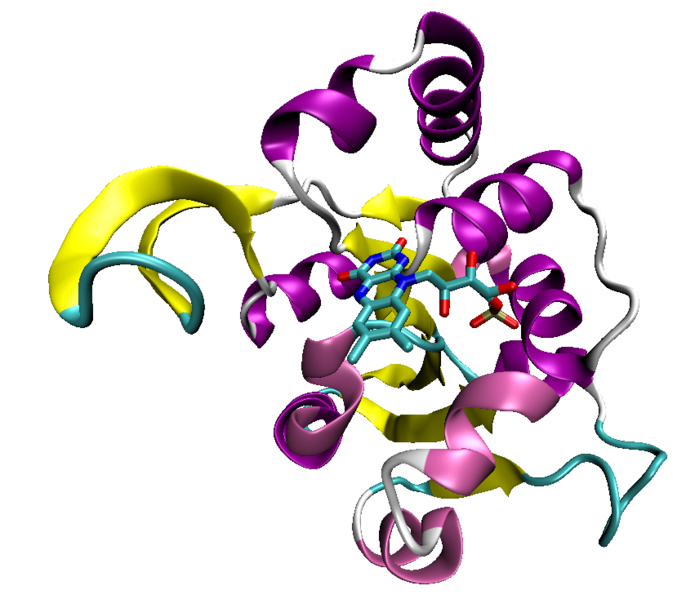
Some proteins that bind ATP do so in a characteristic protein fold known as the Rossmann fold, which is a general nucleotide-binding structural domain that can also bind the cofactor NAD.[30] The most common ATP-binding proteins, known as kinases, share a small number of common folds; the protein kinases, the largest kinase superfamily, all share common structural features specialized for ATP binding and phosphate transfer.[31]
ATP in complexes with proteins generally requires the presence of a divalent cation, almost always magnesium, which binds to the ATP phosphate groups. The presence of magnesium greatly decreases the dissociation constant of ATP from its protein binding partner without affecting the ability of the enzyme to catalyze its reaction once the ATP has bound.[32] The presence of magnesium ions can serve as a mechanism for kinase regulation.[33]
ATP analogs
Biochemistry laboratories often use in vitro studies to explore ATP-dependent molecular processes. Enzyme inhibitors of ATP-dependent enzymes such as kinases are needed to examine the binding sites and transition states involved in ATP-dependent reactions. ATP analogs are also used in X-ray crystallography to determine a protein structure in complex with ATP, often together with other substrates.
Most useful ATP analogs cannot be hydrolyzed as ATP would be; instead they trap the enzyme in a structure closely related to the ATP-bound state. Adenosine 5'-(gamma-thiotriphosphate) is an extremely common ATP analog in which one of the gamma-phosphate oxygens is replaced by a sulfur atom; this molecule is hydrolyzed at a dramatically slower rate than ATP itself and functions as an inhibitor of ATP-dependent processes. In crystallographic studies, hydrolysis transition states are modeled by the bound vanadate ion. However, caution is warranted in interpreting the results of experiments using ATP analogs, since some enzymes can hydrolyze them at appreciable rates at high concentration.[34]
See also
- Adenosine diphosphate (ADP)
- Adenosine monophosphate (AMP)
- Cyclic adenosine monophosphate (cAMP)
- ATPases
- ATP hydrolysis
- Citric acid cycle (also called the Krebs cycle or TCA cycle)
- Phosphagen
- Nucleotide exchange factor
- Mitochondria
- Photophosphorylation
References
- ↑ Knowles JR (1980). "Enzyme-catalyzed phosphoryl transfer reactions". Annu. Rev. Biochem. 49: 877–919. PMID 6250450.
- ↑ Lohmann, K. (1929) Über die Pyrophosphatfraktion im Muskel. Naturwissenschaften 17, 624–625.
- ↑ Lipman F. (1941) Adv. Enzymol. 1, 99-162.
- ↑ Stecher P.G., (1968) The Merck Index 8th edition, Merck and Co. Ltd.
- ↑ Nicholls D.G. and Ferguson S.J. (2002) Bioenergetics Academic press 3rd edition ISBN 0-125-18121-3
- ↑ Gajewski E, Steckler D, Goldberg R (1986). "Thermodynamics of the hydrolysis of adenosine 5'-triphosphate to adenosine 5'-diphosphate" (PDF). J Biol Chem. 261 (27): 12733–7. PMID 3528161.
- ↑ Stryer, Lubert (2002). Biochemistry, fifth edition. New York: W.H. Freeman and Company. ISBN 0-7167-1843-X.
- ↑ 8.0 8.1 Storer A, Cornish-Bowden A (1976). "Concentration of MgATP2− and other ions in solution. Calculation of the true concentrations of species present in mixtures of associating ions". Biochem J. 159 (1): 1–5. PMID 11772.
- ↑ Wilson J, Chin A (1991). "Chelation of divalent cations by ATP, studied by titration calorimetry". Anal Biochem. 193 (1): 16–9. PMID 1645933.
- ↑ Garfinkel L, Altschuld R, Garfinkel D (1986). "Magnesium in cardiac energy metabolism". J Mol Cell Cardiol. 18 (10): 1003–13. PMID 3537318.
- ↑ Beis I., and Newsholme E. A. (1975). The contents of adenine nucleotides, phosphagens and some glycolytic intermediates in resting muscles from vertebrates and invertebrates. Biochem J 152, 23-32.
- ↑ 12.0 12.1 12.2 12.3 Lodish, H, Berk A, Matsudaira P, Kaiser CA, Krieger M, Scott MP, Zipursky SL, Darnell J. (2004). Molecular Cell Biology, 5th, New York: WH Freeman.
- ↑ Parsons M (2004). "Glycosomes: parasites and the divergence of peroxisomal purpose". Mol Microbiol. 53 (3): 717–24. PMID 15255886.
- ↑ 14.0 14.1 14.2 14.3 14.4 14.5 14.6 Voet D, Voet JG. (2004). Biochemistry Vol 1 3rd ed. Wiley: Hoboken, NJ.
- ↑ Abrahams J, Leslie A, Lutter R, Walker J (1994). "Structure at 2.8 A resolution of F1-ATPase from bovine heart mitochondria". Nature. 370 (6491): 621–8. PMID 8065448.
- ↑ 16.0 16.1 Dahout-Gonzalez C, Nury H, Trézéguet V, Lauquin G, Pebay-Peyroula E, Brandolin G. "Molecular, functional, and pathological aspects of the mitochondrial ADP/ATP carrier". Physiology (Bethesda). 21: 242–9. PMID 16868313.
- ↑ Ronnett G, Kim E, Landree L, Tu Y (2005). "Fatty acid metabolism as a target for obesity treatment". Physiol Behav. 85 (1): 25–35. PMID 15878185.
- ↑ Zumft W (1997). "Cell biology and molecular basis of denitrification". Microbiol Mol Biol Rev. 61 (4): 533–616. PMID 9409151.
- ↑ Drake H, Daniel S, Küsel K, Matthies C, Kuhner C, Braus-Stromeyer S (1997). "Acetogenic bacteria: what are the in situ consequences of their diverse metabolic versatilities?". Biofactors. 6 (1): 13–24. PMID 9233536.
- ↑ Allen J (2002). "Photosynthesis of ATP-electrons, proton pumps, rotors, and poise". Cell. 110 (3): 273–6. PMID 12176312.
- ↑ Di Carlo, S. E. and Coliins, H. L. (2001) Advan. Physiol. Edu. 25: 70-71. [1]
- ↑ Joyce CM, Steitz TA (1995). "Polymerase structures and function: variations on a theme?". J. Bacteriol. 177 (22): 6321–9. PMID 7592405.
- ↑ Fields, R.D. and Burnstock G. 2006. Purinergic signalling in neuron-glia interactions. Nature Reviews Neuroscience 7: 423-436.
- ↑ Fredholm, BB, Abbracchio, MP, Burnstock, G, Daly, JW, Harden, TK, Jacobson, KA, Leff, P, Williams, M Nomenclature and classification of purinoceptors Pharmacol Rev 1994 46: 143-156[2]
- ↑ Mishra N, Tuteja R, Tuteja N (2006). "Signaling through MAP kinase networks in plants". Arch Biochem Biophys. 452 (1): 55–68. PMID 16806044.
- ↑ Kamenetsky M, Middelhaufe S, Bank E, Levin L, Buck J, Steegborn C (2006). "Molecular details of cAMP generation in mammalian cells: a tale of two systems". J Mol Biol. 362 (4): 623–39. PMID 16934836.
- ↑ Hanoune J, Defer N. "Regulation and role of adenylyl cyclase isoforms". Annu Rev Pharmacol Toxicol. 41: 145–74. PMID 11264454.
- ↑ 28.0 28.1 Stubbe J (1990). "Ribonucleotide reductases: amazing and confusing". J Biol Chem. 265 (10): 5329–32. PMID 2180924.
- ↑ Chimploy K, Tassotto M, Mathews C (2000). "Ribonucleotide reductase, a possible agent in deoxyribonucleotide pool asymmetries induced by hypoxia". J Biol Chem. 275 (50): 39267–71. PMID 11006282.
- ↑ Rao S, Rossmann M (1973). "Comparison of super-secondary structures in proteins". J Mol Biol. 76 (2): 241–56. PMID 4737475.
- ↑ Scheeff E, Bourne P (2005). "Structural evolution of the protein kinase-like superfamily". PLoS Comput Biol. 1 (5): e49. PMID 16244704.
- ↑ Saylor P, Wang C, Hirai T, Adams J (1998). "A second magnesium ion is critical for ATP binding in the kinase domain of the oncoprotein v-Fps". Biochemistry. 37 (36): 12624–30. PMID 9730835.
- ↑ Lin X, Ayrapetov M, Sun G. "Characterization of the interactions between the active site of a protein tyrosine kinase and a divalent metal activator". BMC Biochem. 6: 25. PMID 16305747.
- ↑ Resetar AM, Chalovich JM. (1995). Adenosine 5'-(gamma-thiotriphosphate): an ATP analog that should be used with caution in muscle contraction studies. 34(49):16039-45.
External links
- CliffNotes - Adenosine Triphosphate (ATP)
- PubChem entry for Adenosine Triphosphate
- KEGG entry for Adenosine Triphosphate
Template:Nucleic acids Template:Enzyme cofactors
ar:أدينوسين ثلاثي الفوسفات be:Адэназінтрыфосфарная кіслата bg:Аденозинтрифосфат ca:Trifosfat d'adenosina cs:Adenosintrifosfát da:Adenosintrifosfat de:Adenosintriphosphat et:Adenosiintrifosfaat gl:ATP ko:아데노신 삼인산 hr:Adenozin trifosfat id:Adenosin trifosfat is:Adenósínþrífosfat it:Adenosintrifosfato he:ATP lb:Adenosintriphosphat lt:ATP hu:Adenozin-trifoszfát mk:Аденозин трифосфат ms:Adenosina trifosfat nl:Adenosinetrifosfaat no:Adenosintrifosfat oc:Adenosina trifosfata ksh:Adenosiin Trifoßfaat simple:Adenosine Triphosphate sk:Adenozíntrifosfát sl:Adenozintrifosfat sr:Аденозин трифосфат sh:Adenozin trifosfat su:Adénosin trifosfat fi:Adenosiinitrifosfaatti sv:Adenosintrifosfat uk:Аденозинтрифосфат