Coronavirus pathophysiology
|
Coronavirus Microchapters |
|
Diagnosis |
|---|
|
Treatment |
|
Case Studies |
|
Coronavirus pathophysiology On the Web |
|
American Roentgen Ray Society Images of Coronavirus pathophysiology |
|
Risk calculators and risk factors for Coronavirus pathophysiology |
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [1]; Associate Editor(s)-in-Chief: Aditya Govindavarjhulla, M.B.B.S. [2]
Pathophysiology
Structure
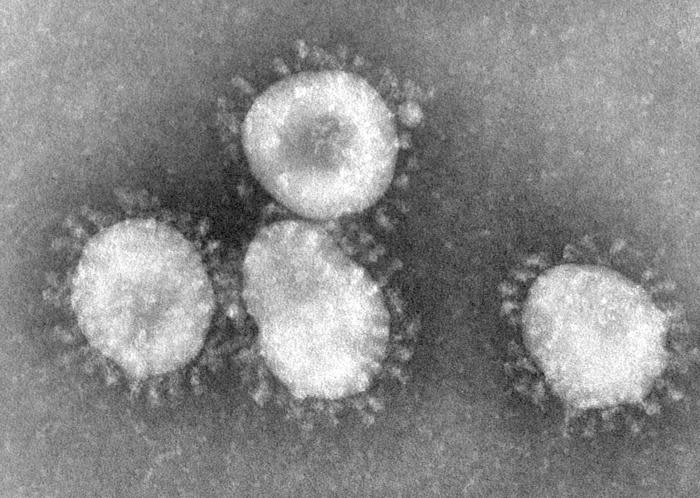
Coronaviruses are enveloped viruses with a positive-sense single-stranded RNA genome and a helical symmetry. The genomic size of coronaviruses ranges from approximately 16 to 31 kilobases, extraordinarily large for an RNA virus. The name coronavirus is derived from the greek (κορώνα, meaning crown) as the virus envelope appears under electron microscopy (E.M.) to be crowned by a characteristic ring of small bulbous structures. This morphology is actually formed by the viral spike (S) peplomers, which are proteins that populate the surface of the virus and determine host tropism. Coronaviruses are grouped in the order Nidovirales, named for the Latin (nidus, meaning nest) as all viruses in this order produce a 3' co-terminal nested set of subgenomic mRNA's during infection.
Proteins that contribute to the overall structure of all coronaviruses are the spike (S), envelope (E), membrane (M) and nucleocapsid (N). In the specific case of SARS, a defined receptor-binding domain on S mediates the attachment of the virus to its cellular receptor, angiotensin-converting enzyme 2 (ACE2).[1] Members of the group 2 coronaviruses also have a shorter spike-like protein called hemagglutinin esterase (HE) encoded in their genome, but for some reason this protein is not always brought to expression (produced) in the cell.[2]
Replication of Coronaviruses
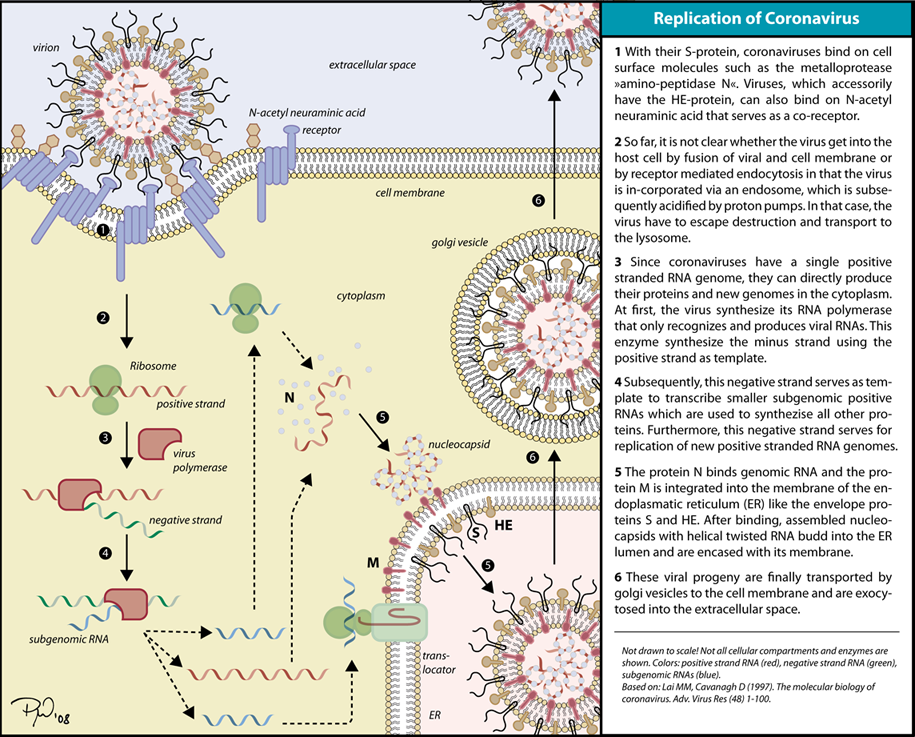
Replication of Coronavirus begins with entry to the cell takes place in the cytoplasm in a membrane-protected microenvironment, upon entry to the cell the virus particle is uncoated and the RNA genome is deposited into the cytoplasm. The Coronavirus genome has a 5’ methylated cap and a 3’polyadenylated-A tail to make it look as much like the host RNA as possible. This also allows the RNA to attach to ribosomes for translation. Coronaviruses also have a protein known as a replicase encoded in its genome which allows the RNA viral genome to be translated into RNA by using the host cells machinery. The replicase is the first protein to be made as once the gene encoding the replicase is translated the translation is stopped by a stop codon. This is known as a nested transcript, where the transcript only encodes one gene- it is monocistronic. The RNA genome is replicated and a long polyprotein is formed, where all of the proteins are attached. Coronaviruses have a non-structural protein called a protease which is able to separate the proteins in the chain. This is a form of genetic economy for the virus allowing it to encode the most amounts of genes in a small amount of nucleotides.
Coronavirus transcription involves a discontinuous RNA synthesis (template switch) during the extension of a negative copy of the subgenomic mRNAs. Base pairing during transcription is a requirement. Coronavirus N protein is required for coronavirus RNA synthesis, and has RNA chaperone activity that may be involved in template switch. Both viral and cellular proteins are required for replication and transcription. Coronaviruses initiate translation by cap-dependent and cap-independent mechanisms. Cell macromolecular synthesis may be controlled after Coronavirus infection by locating some virus proteins in the host cell nucleus. Infection by different coronaviruses cause in the host alteration in the transcription and translation patterns, in the cell cycle, the cytoskeleton, apoptosis and coagulation pathways, inflammation, and immune and stress responses.[3]
Transmission
They are transmitted by aerosols of respiratory secretions, by the fecal-oral route, and by mechanical transmission. Most virus growth occurs in epithelial cells. Occasionally the liver, kidneys, heart or eyes may be infected, as well as other cell types such as macrophages. In cold-type respiratory infections, growth appears to be localized to the epithelium of the upper respiratory tract, but there is no adequate animal model for the human respiratory coronaviruses.
- Incubation period: 2 - 4 days.
- Communicability: Human-to-human transmission is possible during the presence of infectious droplets, which can cause infection via inhalation, or through contaminated surfaces.
- Dissemination
- Reservoir: Humans.
- Vectors: None.
- Dissemination: None.
Role of Spike Proteins
- They induce neutralizing antibody.
- They are important in relating host cell tropism.
- Hemagglutination.
- They mediate the cell to cell or cell to viral fusion by the interaction between viral envelope and the specific receptor of host cell membrane.
Pathogenicity
Coronaviruses primarily infect the upper respiratory and gastrointestinal tract of mammals and birds. Four to five different currently known strains of coronaviruses infect humans. The most publicized human coronavirus, SARS-CoV which causes SARS, has a unique pathogenesis because it causes both upper and lower respiratory tract infections and can also cause gastroenteritis. Coronaviruses are believed to cause a significant percentage of all common colds in human adults. Coronaviruses cause colds in humans primarily in the winter and early spring seasons. The significance and economic impact of coronaviruses as causative agents of the common cold are hard to assess because, unlike rhinoviruses (another common cold virus), human coronaviruses are difficult to grow in the laboratory.
These viruses infect a variety of mammals and birds. The exact number of human isolates are not known as many cannot be grown in culture. In humans, they cause:
- Respiratory infections (common), including Severe Acute Respiratory Syndrome (SARS).
- Enteric infections (occasional - mostly in infants <12 months).
- Neurological syndromes (rare).
Coronaviruses also cause a range of diseases in farm animals and domesticated pets, some of which can be serious and are a threat to the farming industry. Economically significant coronaviruses of farm animals include porcine coronavirus (transmissible gastroenteritis, TGE) and bovine coronavirus, which both result in diarrhea in young animals. Feline enteric coronavirus is a pathogen of minor clinical significance, but spontaneous mutation of this virus can result in feline infectious peritonitis (FIP), a disease associated with high mortality. There are two types of canine coronavirus (CCoV), one that causes mild gastrointestinal disease and one that has been found to cause respiratory disease. Mouse hepatitis virus (MHV) is a coronavirus that causes an epidemic murine illness with high mortality, especially among colonies of laboratory mice. Prior to the discovery of SARS-CoV, MHV had been the best studied coronavirus both in vivo and in vitro as well as at the molecular level. Some strains of MHV cause a progressive demyelinating encephalitis in mice which has been used as a murine model for multiple sclerosis. Significant research efforts have been focused on elucidating the viral pathogenesis of these animal coronaviruses, especially by virologists interested in veterinary and zoonotic diseases.
HCoV-229E and HCoV-OC43 cause the common cold, a self-limiting upper respiratory tract infection. Infection can lead to a number of illnesses such as bronchitis, gastroenteritis, progressive demyelinating encephalitis, diarrhea, peritonitis, nasal obstruction, rhinorrhea, sneezing, sore throat and cough. They can cause more severe lower respiratory tract infection, including pneumonia in infants, elderly and immunocompromised individuals.
HCoV-229E is a common agent if coryza, whereas HCoV-OC43 is generally characterized by sore throats.
HCoV-NL63 causes laryngotracheitis (croup) and nonfatal upper and lower respiratory tract infections in children, elderly, and immunocompromised individuals. HCoV-HKU1 causes mild upper respiratory diseases, the common cold, bronchiolitis, and pneumonia, with symptoms such as rhinorrhea, fever, cough, febrile seizure, and wheezing. More severe illness may occur in children, adults with underlying disease, the elderly, and may be associated with gastrointestinal illness.
Laboratory Hazards
No infections have been reported. However, this may be an underestimate of the number of incidences as symptoms are nonspecific and self limiting.
- Source: Specimens of upper and lower respiratory tract.
- Primary Hazard: Aerosols and contact with stools.
Stablity and Viability
- Drug Susceptibility: Currently, there are no specific antiviral drugs for coronavirus available.[4]
- Susceptibility to Disinfectants: Susceptible to 0.1% sodium hypochlorite, 0.1% organochlorine, 10% iodophore, 70% ethanol and 2% glutaraldehyde. Resistant to 0.04% quaternary ammonium compound and phenolics.
- Physical Inactivation: Inactivation by UV light can be done by exposure to 1200 µJ/cm2 for 30 minutes.
- Survival Outside Host: Survives up to six days in aqueous mediums and up to 3 hours on dry inanimate surfaces.
Associated Conditions
Conditions associated with coronavirus infection may include:
- Diabetes[5]
- Cancer
- Chronic lung disease
- Chronic heart disease
- Chronic kidney disease
- Neurologic disease[6]
References
- ↑ Li, Fang, et. al. (2005). "Structure of SARS Coronavirus Spike Receptor-Binding Domain Complexed with Receptor". Science. 309: 1864&ndash, 1868. doi:10.1126/science.1116480.
- ↑ de Haan CAM, Rottier PJM (2005). "Molecular Interactions in the Assembly of Coronaviruses". Advances in Virus Research. 64: 185&ndash, 186.
- ↑ Enjuanes; et al. (2008). "Coronavirus Replication and Interaction with Host". Animal Viruses: Molecular Biology. Caister Academic Press. ISBN 978-1-904455-22-6.
- ↑ https://www.cdc.gov/coronavirus/mers/about/symptoms.html. Missing or empty
|title=(help) - ↑ Arabi YM, Harthi A, Hussein J, Bouchama A, Johani S, Hajeer AH, Saeed BT, Wahbi A, Saedy A, AlDabbagh T, Okaili R, Sadat M, Balkhy H (August 2015). "Severe neurologic syndrome associated with Middle East respiratory syndrome corona virus (MERS-CoV)". Infection. 43 (4): 495–501. doi:10.1007/s15010-015-0720-y. PMC 4521086. PMID 25600929.