ATP synthase
|
WikiDoc Resources for ATP synthase |
|
Articles |
|---|
|
Most recent articles on ATP synthase Most cited articles on ATP synthase |
|
Media |
|
Powerpoint slides on ATP synthase |
|
Evidence Based Medicine |
|
Clinical Trials |
|
Ongoing Trials on ATP synthase at Clinical Trials.gov Clinical Trials on ATP synthase at Google
|
|
Guidelines / Policies / Govt |
|
US National Guidelines Clearinghouse on ATP synthase
|
|
Books |
|
News |
|
Commentary |
|
Definitions |
|
Patient Resources / Community |
|
Patient resources on ATP synthase Discussion groups on ATP synthase Patient Handouts on ATP synthase Directions to Hospitals Treating ATP synthase Risk calculators and risk factors for ATP synthase
|
|
Healthcare Provider Resources |
|
Causes & Risk Factors for ATP synthase |
|
Continuing Medical Education (CME) |
|
International |
|
|
|
Business |
|
Experimental / Informatics |
Overview
An ATP synthase (EC 3.6.3.14) is a general term for an enzyme that can synthesize adenosine triphosphate (ATP) from adenosine diphosphate (ADP) and inorganic phosphate by using some form of energy. This energy is often in the form of protons moving down a electrochemical gradient, such as from the lumen into the stroma of chloroplasts or from the inter-membrane space into the matrix in mitochondria. The overall reaction sequence is:
- ADP + Pi → ATP
These enzymes are of crucial importance in almost all organisms, because ATP is the common "energy currency" of cells.
The antibiotic oligomycin inhibits the FO unit of ATP synthase.
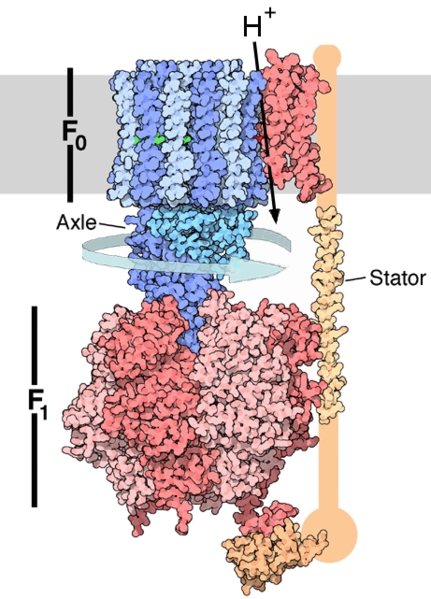
Structure and nomenclature
In mitochondria, the F1FO ATP synthase has a long history of scientific study.
- the FO portion is within the membrane.
- The F1 portion of the ATP synthase is above the membrane.
It's easy to visualize the FOF1 particle as resembling the fruiting body of a common mushroom, with the head being the F1 particle, the stalk being the gamma subunit of F1, and the base and "roots" being the FO particle embedded in the membrane.
The nomenclature of the enzyme suffers from a long history. The F1 fraction derives it name from the term "Fraction 1" and FO (written as a subscript "O", not "zero") derives its name from being the oligomycin binding fraction.
Taking as an example the nomenclature of subunits in the bovine enzyme, many subunits have alphabet names:
- Greek letters: alpha, beta, gamma, delta, epsilon
- Roman letters: a, b, c, d, e, f, g, h
Others have more complex names:
- F6 (from "Fraction 6")
- OSCP (the oligomycin sensitivity conferral protein), ATP5O
- A6L (named for the gene that codes for it in the mitochondrial genome)
- IF1 (inhibitory factor 1), ATPIF1
The F1 particle is large and can be seen in the transmission electron microscope by negative staining.[1] These are particles of 9 nm diameter that pepper the inner mitochondrial membrane. They were originally called elementary particles and were thought to contain the entire respiratory apparatus of the mitochondrion, but through a long series of experiments, Ephraim Racker and his colleagues (who first isolated the F1 particle in 1961) were able to show that this particle is correlated with ATPase activity in uncoupled mitochondria and with the ATPase activity in submitochondrial particles created by exposing mitochondria to ultrasound. This ATPase activity was further associated with the creation of ATP by a long series of experiments in many laboratories.
Binding change mechanism
In the 1960s through the 1970s, Paul Boyer developed his binding change, or flip-flop, mechanism, which postulated that ATP synthesis is coupled with a conformational change in the ATP synthase generated by rotation of the gamma subunit. The research group of John E. Walker, then at the MRC Laboratory of Molecular Biology in Cambridge but now at the MRC Dunn Human Nutrition Unit (also in Cambridge) crystallized the F1 catalytic-domain of ATP synthase. The structure, at the time the largest asymmetric protein structure known, indicated that Boyer's rotary-catalysis model was essentially correct. For elucidating this Boyer and Walker shared half of the 1997 Nobel Prize in Chemistry. Jens Christian Skou received the other half of the Chemistry prize that year "for the first discovery of an ion-transporting enzyme, Na+, K+ -ATPase"

The crystal structure of the F1 showed alternating alpha and beta subunits (3 of each), arranged like segments of an orange around an asymmetrical gamma subunit. According to the current model of ATP synthesis (known as the alternating catalytic model), the proton-motive force across the inner mitochondrial membrane, generated by the electron transport chain, drives the passage of protons through the membrane via the FO region of ATP synthase. A portion of the FO (the ring of c-subunits) rotates as the protons pass through the membrane. The c-ring is tightly attached to the asymmetric central stalk (consisting primarily of the gamma subunit) which rotates within the alpha3beta3 of F1 causing the 3 catalytic nucleotide binding sites to go through a series of conformational changes that leads to ATP synthesis. The major F1 subunits are prevented from rotating in sympathy with the central stalk rotor by a peripheral stalk that joins the alpha3beta3 to the non-rotating portion of FO. The structure of the intact ATP synthase is currently known at low-resolution from electron cryo-microscopy (cryo-EM) studies of the complex. The cryo-EM model of ATP synthase shows that the peripheral stalk is a flexible rope-like structure that wraps around the complex as it joins F1 to FO. Under the right conditions, the enzyme reaction can also be carried out in reverse, with ATP hydrolysis driving proton pumping across the membrane.
The binding change mechanism involves the active site of a β subunit cycling between three states.[2] In the "open" state, ADP and phosphate enter the active site, in the diagram to the right this is shown in brown. The protein then closes up around the molecules and binds them loosely - the "loose" state (shown in red). The enzyme then undergoes another change in shape and forces these molecules together, with the active site in the resulting "tight" state (shown in pink) binding the newly-produced ATP molecule with very high affinity. Finally, the active site cycles back to the open state, releasing ATP and binding more ADP and phosphate, ready for the next cycle of ATP production.
Physiological role
Like other enzymes, the activity of F1FO ATP synthase is reversible. Large enough quantities of ATP cause it to create a transmembrane proton gradient, this is used by fermenting bacteria which do not have an electron transport chain, and hydrolyze ATP to make a proton gradient, which they use for flagella and transport of nutrients into the cell.
In respiring bacteria under physiological conditions, ATP synthase generally runs in the opposite direction, creating ATP while using the protonmotive force created by the electron transport chain as a source of energy. The overall process of creating energy in this fashion is termed oxidative phosphorylation. The same process takes place in mitochondria, where ATP synthase is located in the inner mitochondrial membrane (so that F1-part sticks into mitochondrial matrix, where ATP synthesis takes place).
ATP synthase in different organisms
Plant ATP synthase
In plants ATP synthase is also present in chloroplasts (CF1FO-ATP synthase). The enzyme is integrated into thylakoid membrane; the CF1-part sticks into stroma, where dark reactions of photosynthesis (Also called the light-independent reactions or the Calvin cycle) and ATP synthesis take place. The overall structure and the catalytic mechanism of the chloroplast ATP synthase are almost the same as those of the mitochondrial enzyme. However, in chloroplasts the proton motive force is generated not by respiratory electron transport chain, but by primary photosynthetic proteins.
E. coli ATP synthase
E. coli ATP synthase is the simplest known form of ATP synthase, with 8 different subunit types.
Yeast ATP synthase
Yeast ATP synthase is the most complex known and is made of 20 different types of subunits.
Evolution of ATP synthase
The evolution of ATP synthase is thought to be an example of modular evolution, where two subunits with their own functions have become associated and gained new functionality. The F1 particle shows significant similarity to hexameric DNA helicases and the FO particle shows some similarity to H+ powered flagellar motor complexes.
The α3β3 hexamer of the F1 particle shows significant structural similarity to hexameric DNA helicases; both form a ring with 3 fold rotational symmetry with a central pore. Both also have roles dependent on the relative rotation of a macromolecule within the pore; the DNA helicases use the helical shape of DNA to drive their motion along the DNA molecule and to detect supercoiling whilst the α3β3 hexamer uses the conformational changes due rotation of the γ subunit to drive an enzymatic reaction.
The H+ motor of the FO particle shows great functional similarity to the H+ motors seen in flagellar motors. Both feature a ring of many small alpha helical proteins which rotate relative to nearby stationary proteins using a H+ potential gradient as an energy source. This is, however, a fairly tenuous link - the overall structure of flagellar motors is far more complex than the FO particle and the ring of rotating proteins is far larger, with around 30 compared to the 10, 11 or 14 known in the FO complex.
The modular evolution theory for the origin of ATP synthase suggests that two subunits with independent function, a DNA helicase with ATPase activity and a H+ motor, were able to bind, and the rotation of the motor drive the ATPase activity of the helicase in reverse. This would then evolve to become more efficient, and eventually develop into the complex ATP synthases seen today. Alternatively the DNA helicase/H+ motor complex may have had H+ pump activity, the ATPase activity of the helicase driving the H+ motor in reverse. This could later evolve to carry out the reverse reaction and act as an ATP synthase.
See also
- Oxidative phosphorylation
- Mitochondrion
- Chloroplast
- Electron transfer chain
- Proton pump
- Transmembrane ATPase
- Flavoprotein
- ATP synthase subunit C
References
- ↑ Fernandez-Moran et al., Journal of Molecular Biology, Vol 22, p 63, 1962
- ↑ Gresser MJ, Myers JA, Boyer PD (1982). "Catalytic site cooperativity of beef heart mitochondrial F1 adenosine triphosphatase. Correlations of initial velocity, bound intermediate, and oxygen exchange measurements with an alternating three-site model". J. Biol. Chem. 257 (20): 12030–8. PMID 6214554.
External links
- "ATP synthase - a splendid molecular machine" at University of Osnabrück
- Well illustrated ATP synthase lecture by Antony Crofts of the University of Illinois at Urbana-Champaign.
- Proton and Sodium translocating F-type, V-type and A-type ATPases in OPM database
- The Nobel Prize in Chemistry 1997 to Paul D. Boyer and John E. Walker for the enzymatic mechanism of synthesis of ATP; and to Jens C. Skou, for discovery of an ion-transporting enzyme, Na+, K+-ATPase.
de:ATP-Synthase it:ATPasi trasportante H+ tra due settori su:ATP sintase sv:ATP-syntas