CRE box gene transcription laboratory
Editor-In-Chief: Henry A. Hoff
{{free media}}A laboratory is a specialized activity, a construct, you create where you as a student, teacher, or researcher can have hands-on, or as close to hands-on as possible, experience actively analyzing an entity, source, or object of interest. Usually, there's more to do than just analyzing. The construct is often a room, building or institution equipped for scientific research, experimentation as well as analysis.
This laboratory is a continuation of the previous laboratory.
In the room next door is an astronaut on the Mars expedition, three months along on the six-month journey. A physician and lab assistants have been performing tests on her. Because she has been in zero gravity for more than three months her body chemistry and anatomy now differ from what it was in the controlled gravity environment of Earth. She has lost about 10 % each of her bone, muscle, and brain mass. Comparisons with gene expression sequences now and when on Earth have found that the gene expression for alpha-1-B glycoprotein is not normal. If a way to correct this expression cannot be found she must be returned to Earth maybe to recover, maybe not!
But, it is unlikely she will survive three more months at zero g either to be returned to Earth or put on Mars. Worse, the microgravity may not be the only culprit. There is also the radiation of the interplanetary medium.
You have been tasked to examine her DNA to confirm, especially with the extended data between ZNF497 and A1BG, the presence or absence of CRE boxes regarding the possible expression of alpha-1-B glycoprotein.
The CRE boxes are gene transcription factors (TF).
CRE boxes
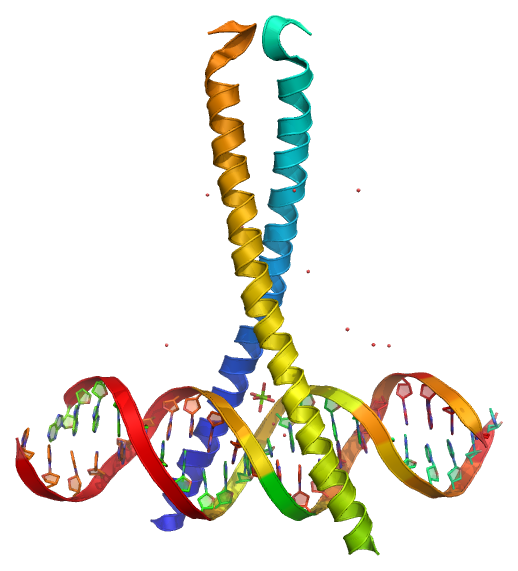
CREB (cAMP response element-binding protein)[1] is a cellular transcription factor that binds to certain DNA sequences called cAMP response elements (CRE), thereby increasing or decreasing the transcription of the genes.[2] CREB was first described in 1987 as a cyclic adenosine monophosphate (cAMP)-responsive transcription factor regulating the somatostatin gene.[3]
Genes whose transcription is regulated by CREB include: c-fos, BDNF, tyrosine hydroxylase, numerous neuropeptides (such as somatostatin, enkephalin, VGF, corticotropin-releasing hormone),[2] and genes involved in the mammalian circadian clock (PER1, PER2).[4]
CREB has a well-documented role in neuronal plasticity and long-term memory formation in the brain and has been shown to be integral in the formation of spatial memory.[5] CREB downregulation is implicated in the pathology of Alzheimer's disease and increasing the expression of CREB is being considered as a possible therapeutic target for Alzheimer's disease.[6]
Consensus sequence
"Each species' promoter contains a cAMP/response element (CRE)1 consensus sequence ([7]) upstream of a TATA box. Two other members of this family of proteins, chromogranin B and secretogranin II (also known as chromogranin C), contain similar CRE and TATA homologies in their proximal gene promoters (8, 9)."[8]
"Within the cAMP-responsive element of the somatostatin gene, we observed an 8-base palindrome, 5'-TGACGTCA-3', which is highly conserved in many other genes whose expression is regulated by cAMP."[7]
cAMP response element
The cAMP response element (CRE) is the response element for CREB which contains the highly conserved nucleotide sequence, 5'-TGACGTCA-3’. CRE sites are typically found upstream of genes, within the promoter or enhancer regions.[9] There are approximately 750,000 palindromic and half-site CREs in the human genome. However, the majority of these sites remain unbound due to cytosine methylation.[10]
Nucleotides
DNA mapping has been performed. Her DNA for A1BG promoters can be found at A1BG gene transcriptions#Nucleotides.
Programming
Sample programs for preparing test programs are available at Gene transcriptions/A1BG/Programming.
Hypotheses
- CRE boxes are not involved in the transcription of A1BG.
Core promoters
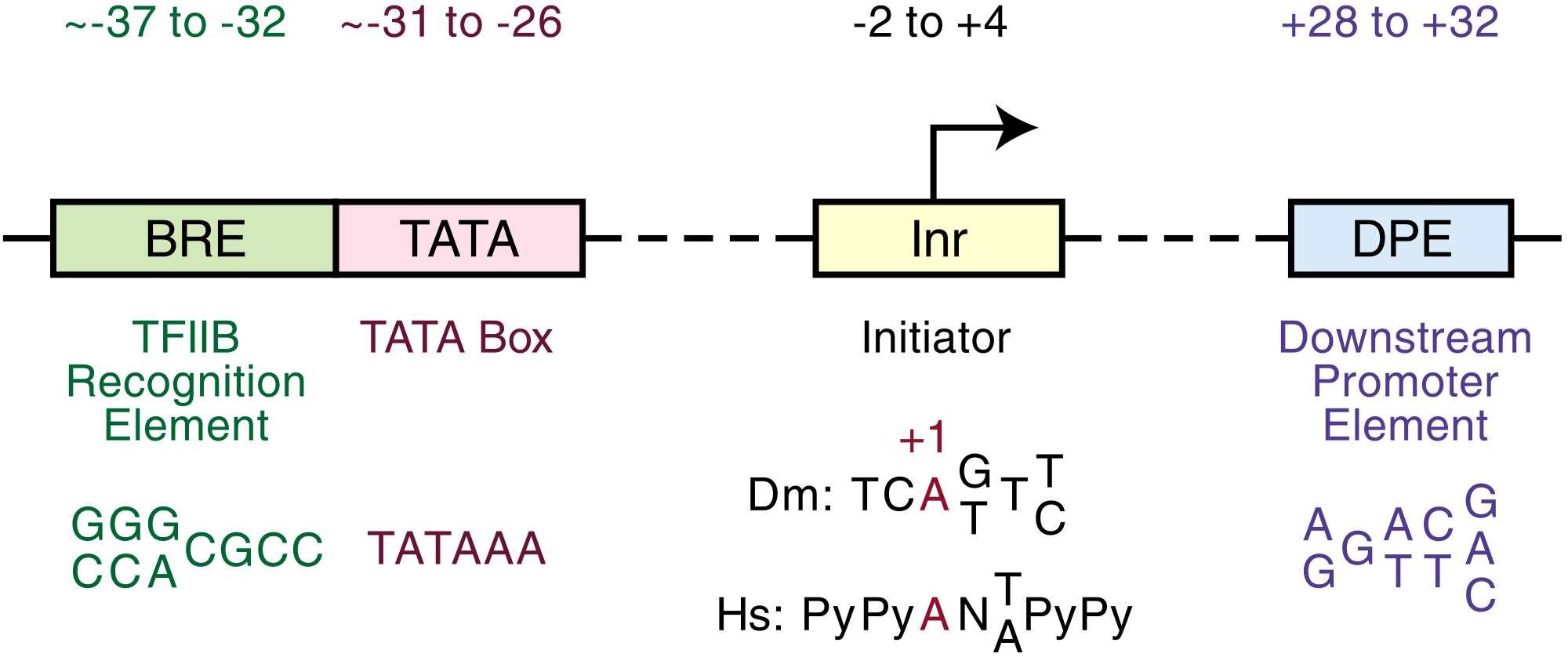
The core promoter is approximately -34 nts upstream from the TSS.
From the first nucleotide just after ZSCAN22 to the first nucleotide just before A1BG are 4460 nucleotides. The core promoter on this side of A1BG extends from approximately 4425 to the possible transcription start site at nucleotide number 4460.
To extend the analysis from inside and just on the other side of ZNF497 some 3340 nts have been added to the data. This would place the core promoter some 3340 nts further away from the other side of ZNF497. The TSS would be at about 4300 nts with the core promoter starting at 4266.
Def. "the factors, including RNA polymerase II itself, that are minimally essential for transcription in vitro from an isolated core promoter" is called the basal machinery, or basal transcription machinery.[12]
Proximal promoters
Def. a "promoter region [juxtaposed to the core promoter that] binds transcription factors that modify the affinity of the core promoter for RNA polymerase.[12][13]"[13] is called a proximal promoter.
The proximal sequence upstream of the gene that tends to contain primary regulatory elements is a proximal promoter.
It is approximately 250 base pairs or nucleotides, nts, upstream of the transcription start site.
The proximal promoter begins about nucleotide number 4210 in the negative direction.
The proximal promoter begins about nucleotide number 4195 in the positive direction.
Distal promoters
The "upstream regions of the human [cytochrome P450 family 11 subfamily A] CYP11A and bovine CYP11B genes [have] a distal promoter in each gene. The distal promoters are located at −1.8 to −1.5 kb in the upstream region of the CYP11A gene and −1.5 to −1.1 kb in the upstream region of the CYP11B gene."[14]
"Using cloned chicken βA-globin genes, either individually or within the natural chromosomal locus, enhancer-dependent transcription is achieved in vitro at a distance of 2 kb with developmentally staged erythroid extracts. This occurs by promoter derepression and is critically dependent upon DNA topology. In the presence of the enhancer, genes must exist in a supercoiled conformation to be actively transcribed, whereas relaxed or linear templates are inactive. Distal protein–protein interactions in vitro may be favored on supercoiled DNA because of topological constraints."[15]
Distal promoter regions may be a relatively small number of nucleotides, fairly close to the TSS such as (-253 to -54)[16] or several regions of different lengths, many nucleotides away, such as (-2732 to -2600) and (-2830 to -2800).[17]
The "[d]istal promoter is not a spacer element."[18]
Using an estimate of 2 knts, a distal promoter to A1BG would be expected after nucleotide number 2460.
Any transcription factor before A1BG from the direction of ZN497 may be out to 2300 nts.
Samplings
Regarding hypothesis 1
For the Basic programs (starting with SuccessablesCRE.bas) written to compare nucleotide sequences with the sequences on either the template strand (-), or coding strand (+), of the DNA, in the negative direction (-), or the positive direction (+), including extension of nts from 958 to 4445, the programs are, are looking for, and found:
- negative strand in the negative direction (from ZSCAN22 to A1BG) is SuccessablesCRE--.bas, looking for 3'-TGACGTCA-5', 1, 3'-TGACGTCA-5', 4317,
- negative strand in the positive direction (from ZNF497 to A1BG) is SuccessablesCRE-+.bas, looking for 3'-TGACGTCA-5', 0,
- positive strand in the negative direction is SuccessablesCRE+-.bas, looking for 3'-TGACGTCA-5', 0,
- positive strand in the positive direction is SuccessablesCRE++.bas, looking for 3'-TGACGTCA-5', 0,
- complement, negative strand, negative direction is SuccessablesCREc--.bas, looking for 3'-ACTGCAGT-5', 0,
- complement, negative strand, positive direction is SuccessablesCREc-+.bas, looking for 3'-ACTGCAGT-5', 0,
- complement, positive strand, negative direction is SuccessablesCREc+-.bas, looking for 3'-ACTGCAGT-5', 1, 3'-ACTGCAGT-5', 4317,
- complement, positive strand, positive direction is SuccessablesCREc++.bas, looking for 3'-ACTGCAGT-5', 0,
- inverse complement, negative strand, negative direction is SuccessablesCREci--.bas, looking for 3'-TGACGTCA-5', 1, 3'-TGACGTCA-5', 4317,
- inverse complement, negative strand, positive direction is SuccessablesCREci-+.bas, looking for 3'-TGACGTCA-5', 0,
- inverse complement, positive strand, negative direction is SuccessablesCREci+-.bas, looking for 3'-TGACGTCA-5', 0,
- inverse complement, positive strand, positive direction is SuccessablesCREci++.bas, looking for 3'-TGACGTCA-5', 0,
- inverse, negative strand, negative direction, is SuccessablesCREi--.bas, looking for 3'-ACTGCAGT-5', 0,
- inverse, negative strand, positive direction, is SuccessablesCREi-+.bas, looking for 3'-ACTGCAGT-5', 0,
- inverse, positive strand, negative direction, is SuccessablesCREi+-.bas, looking for 3'-ACTGCAGT-5', 1, 3'-ACTGCAGT-5', 4317,
- inverse, positive strand, positive direction, is SuccessablesCREi++.bas, looking for 3'-ACTGCAGT-5', 0.
Verifications
To verify that your sampling has explored something, you may need a control group. Perhaps where, when, or without your entity, source, or object may serve.
Another verifier is reproducibility. Can you replicate something about your entity in your laboratory more than 3 times. Five times is usually a beginning number to provide statistics (data) about it.
For an apparent one time or perception event, document or record as much information coincident as possible. Was there a butterfly nearby?
Has anyone else perceived the entity and recorded something about it?
Gene ID: 1, includes the nucleotides between neighboring genes and A1BG. These nucleotides can be loaded into files from either gene toward A1BG, and from template and coding strands. These nucleotide sequences can be found in Gene transcriptions/A1BG. Copying the above discovered CRE boxes and putting the sequences in "⌘F" locates these sequences in the same nucleotide positions as found by the computer programs.
Core promoters CRE boxes
From the first nucleotide just after ZSCAN22 to the first nucleotide just before A1BG are 4460 nucleotides. The core promoter on this side of A1BG extends from approximately 4425 to the possible transcription start site at nucleotide number 4460.
On the negative strand in the negative direction (from ZSCAN22 to A1BG), looking for 3'-TGACGTCA-5', there no CRE boxes in the core promoter.
From the first nucleotide just after ZNF497 to the first nucleotide just before A1BG are 858 nucleotides. The core promoter on this side of A1BG extends from approximately 824 to the possible transcription start site at nucleotide number 858. Nucleotides (nts) have been added from ZNF497 to A1BG. The TSS for A1BG is now at 4300 nts from just on the other side of ZNF497. The core promoter should now be from 4266 to 4300.
On the negative strand in the positive direction (from ZNF497 to A1BG), looking for 3'-TGACGTCA-5', there no CRE boxes in the core promoter.
Proximal promoter CRE boxes
The proximal promoter begins about nucleotide number 4210 in the negative direction.
There is one CRE box in the negative direction at 4317.
The proximal promoter begins about nucleotide number 4195 in the positive direction.
There are no CRE boxes in the positive direction.
Distal promoter CRE boxes
Using an estimate of 2 knts, a distal promoter to A1BG would be expected after nucleotide number 2460.
There are no CRE boxes after nucleotide number 2460 in the distal promoter.
There are no CRE boxes after nucleotide number 2300.
Transcribed CRE boxes
GeneID: 627 BDNF brain derived neurotrophic factor. Also known as ANON2; BULN2. "This gene encodes a member of the nerve growth factor family of proteins. Alternative splicing results in multiple transcript variants, at least one of which encodes a preproprotein that is proteolytically processed to generate the mature protein. Binding of this protein to its cognate receptor promotes neuronal survival in the adult brain. Expression of this gene is reduced in Alzheimer's, Parkinson's, and Huntington's disease patients. This gene may play a role in the regulation of the stress response and in the biology of mood disorders."[19]
GeneID: 1113 CHGA chromogranin A. Also known as CGA. "The protein encoded by this gene is a member of the chromogranin/secretogranin family of neuroendocrine secretory proteins. It is found in secretory vesicles of neurons and endocrine cells. This gene product is a precursor to three biologically active peptides; vasostatin, pancreastatin, and parastatin. These peptides act as autocrine or paracrine negative modulators of the neuroendocrine system. Two other peptides, catestatin and chromofungin, have antimicrobial activity and antifungal activity, respectively. Two transcript variants encoding different isoforms have been found for this gene."[20]
Secretory "stimulation of pheochromocytoma cells also activates the biosynthesis of the major secreted protein (chromogranin A), that the activation is transcriptional, and that a small proximal domain, including the CRE box, is, at least in part, both necessary and sufficient to account for the positive response to nicotine."[21]
"A CRE box containing somatostatin promoter (-71 bp to +55 bp)/luciferase reporter plasmid was obtained from Marc Montminy, Salk Institute, La Jolla, CA. Its CRE box (TGACGTCA) was at -41 to -48 bp."[21]
"Since the proximal positive response region [...] contains a CRE homology ((-71 bp)-TGACGTAA-(-64 bp)), we tested the effect of a CRE box point-gap mutation (TGA-GTAA) in a transfected 77-bp chromogranin A promoter/luciferase reporter construct [...]. The nicotinic induction ratio (nicotine stimulation/mock stimulation) fell from 2.58-fold in the wild-type promoter down to 1.64-fold (i.e. a 60% fall, p < 0.05) in the promoter with the CRE box point-gap mutation."[21]
"We then tested whether a CRE box alone could transfer nicotinic responsiveness to a heterologous (TK) promoter (in pTK-LUC). The chromogranin A CRE (TGACGTAA) insert conferred 3.97-fold nicotinic induction [...], while a consensus CRE (TGACGTCA) was induced 4.94-fold by nicotine; each of these nicotinic responses was significantly (p < 0.05) greater than that of the original TK promoter vector [...]. The nicotinic induction was blunted (down to 1.31-fold) by point-gap mutation of the CRE box (TGA-GTAA). Even a CRE box (TGACGTCA) in the transfected promoter of a gene (somatostatin) not ordinarily expressed in chromaffin cells responded 2.03-fold (p < 0.05) to nicotine (from 134 ± 10.1 to 272 ± 19.6 luciferase light units/mg protein; n = 4 transfected wells/condition)."[21]
"In electrophoretic gel mobility shift assays, we tested the effect of PC12 nicotine exposure (at a dose (10-3 M) and time (16 h) which stimulated the transfected promoter [...] on nuclear protein binding to four promoter regions: three that responded positively to nicotine during transfection (-432 to -312, -147 to -77, and -71 to -64 bp (the CRE box)), and one that seemed to respond negatively (-181 to -147 bp). None of these regions showed any significant change in band retardation pattern after nicotine [...]."[21]
"Because CREB-bound CRE boxes are only transcriptionally transactivated when CREB itself has been activated by phosphorylation at serine 133 (33), we undertook antibody supershift assays [...], using not only an antibody directed against CREB (epitope: the CREB kinase-inducible domain), but also an antibody which specifically recognized CREB only in its activated, phosphorylated form (pS133-CREB). Cell exposure to nicotine did not alter the anti-CREB supershift by PC12 nuclear proteins, but did result in a qualitatively new anti-pS133-CREB supershift band. Competition experiments with a 100-fold molar excess of unlabeled CRE [...] indicated that both the anti-CREB and the anti-pS133-CREB supershifts represented CRE-binding complexes."[21]
"The CRE box is at promoter position (-71 bp)TGACGTAA(-64 bp)."[21]
"Each species' promoter contains a cAMP/response element (CRE)1 consensus sequence ([7]) upstream of a TATA box. Two other members of this family of proteins, chromogranin B and secretogranin II (also known as chromogranin C), contain similar CRE and TATA homologies in their proximal gene promoters (8, 9)."[8]
"1. Abbreviations used: CMV, cytomegalovirus; CRE, cAMP responsive element; CREB, CRE-binding protein; CREM, CRE modulator protein; DBH, dopamine ,/-hydroxylase; LUC, luciferase; TK, thymidine kinase."[8]
"Within the cAMP-responsive element of the somatostatin gene, we observed an 8-base palindrome, 5'-TGACGTCA-3', which is highly conserved in many other genes whose expression is regulated by cAMP."[7]
GeneID: 1114 CHGB chromogranin B. Also known as SCG1. "This gene encodes a tyrosine-sulfated secretory protein abundant in peptidergic endocrine cells and neurons. This protein may serve as a precursor for regulatory peptides."[22]
The "isolated mouse chromogranin B promoter [specifically] the proximal chromogranin B promoter (from −216 to −91 bp); [...] contains an E box (at [−206 bp]CACCTG[−201 bp]), four G/C-rich regions (at[− 196 bp]CCCCGC[−191 bp], [−134 bp]CCGCCCGC[−127 bp],[− 125 bp]GGCGCCGCC[−117 bp], and [−115 bp]CGGGGC[−110 bp]), and a cAMP response element (CRE; at [−102 bp]TGACGTCA[−95 bp]). A 60-bp core promoter region, defined by an internal deletion from −134 to −74 bp upstream of the cap site and spanning the CRE and three G/C-rich regions, directed tissue-specific expression of the gene. The CRE motif directed cell type-specific expression of the chromogranin B gene in neurons, whereas three of the G/C-rich regions played a crucial role in neuroendocrine cells. Both the endogenous chromogranin B gene and the transfected chromogranin B promoter were induced by preganglionic secretory stimuli (pituitary adenylyl cyclase-activating polypeptide, vasoactive intestinal peptide, or a nicotinic cholinergic agonist), establishing stimulus-transcription coupling for this promoter. The adenylyl cyclase activator forskolin, nerve growth factor, and retinoic acid also activated the chromogranin B gene. Secretagogue-inducible expression of chromogranin B also mapped onto the proximal promoter; inducible expression was entirely lost upon internal deletion of the 60-bp core (from −134 to −74 bp). [...] CRE and G/C-rich domains are crucial determinants of both cell type-specific and secretagogue-inducible expression of the chromogranin B gene."[23]
"Sequence analysis of 2908 bp of the 5′-flanking region of the mouse chromogranin B gene revealed several consensus matches for cis-acting transcriptional control elements [...] reported as base pairs upstream of the cap site (+1): 1) the TATA box at [−33 bp]TTCATAA[−27 bp]; 2) a cAMP response element (CRE) (24) at [−102 bp]TGACGTCA[−95 bp]; 3) eight G/C-rich elements of 6 consecutive bp or more (potential binding sites for such factors as Sp1, Ap2, or Egr1) at −8/−2 bp, −45/−39 bp, −62/−53 bp, −83/−64 bp, −115/−110 bp, −125/−117 bp, −134/−127 bp, and− 196/−191 bp [one of these G/C-rich sites contains a perfect (9/9 bp) consensus match for an Egr1 recognition site: −79/−71 bp (25); another G/C-rich site overlaps a perfect (10/10 bp) consensus match (GGGRNNYYCC; IUPAC nomenclature) for a nuclear factor-κB site: [−83 bp]GGGGCGCCCC[−74 bp] (25); several of these G/C-rich sites overlap partial matches for Ap2 motifs (e.g. GSSWGSCC; IUPAC nomenclature) (25)]; and 4) an E-box (CANNTG; IUPAC nomenclature) at [−206 bp]CACCTG[−201 bp] (25)."[23]
"Mouse chromogranin B promoter sequence. Nucleotides are numbered from the transcription initiation site (+1) of the gene. Note the positions of the TATA box at [−33 bp]TTCATAA[−27 bp], the CRE at [−102 bp]TGACGTCA[−95 bp], and eight G/C-rich elements of 6 or more consecutive bp (potential binding sites for such factors as Sp1, Ap2, or Egr1) at [−8 bp]CCGCGCC[−2 bp], [−45 bp]CCCCGCC[−39 bp], [−62 bp]CGCCCCCGGG[−53 bp],[−83 bp]GGGGCGCCCCCGCCCGCCGC[−64 bp], [−115bp]CGGGGC[−110 bp], [−125 bp]GGCGCCGCC[−117 bp], [−134]CCGCCCGC[−127 bp], and [−196 bp]CCCCGC[−191 bp]. One of these G/C-rich sites is a perfect (9/9 bp) consensus match for an Egr1 recognition site: [−79 bp]CGCCCCCGC[−71 bp]. One of these G/C-rich sites overlaps a perfect (10/10 bp) consensus match (GGGRNNYYCC; IUPAC nomenclature) for a nuclear factor-κB site: [−83 bp]GGGGCGCCCC[−74 bp]. Several of these G/C-rich sites overlap partial matches for Ap2 motifs (e.g. GSSWGSCC; IUPAC nomenclature). There is an E box (CANNTG; IUPAC nomenclature) at [−206 bp]CACCTG[−201 bp]."[23]
GeneID: 1392 CRH corticotropin releasing hormone. Also known as CRF; CRH1. "This gene encodes a member of the corticotropin-releasing factor family. The encoded preproprotein is proteolytically processed to generate the mature neuropeptide hormone. In response to stress, this hormone is secreted by the paraventricular nucleus (PVN) of the hypothalamus, binds to corticotropin releasing hormone receptors and stimulates the release of adrenocorticotropic hormone from the pituitary gland. Marked reduction in this protein has been observed in association with Alzheimer's disease. Autosomal recessive hypothalamic corticotropin deficiency has multiple and potentially fatal metabolic consequences including hypoglycemia and hepatitis. In addition to production in the hypothalamus, this protein is also synthesized in peripheral tissues, such as T lymphocytes, and is highly expressed in the placenta. In the placenta it is a marker that determines the length of gestation and the timing of parturition and delivery. A rapid increase in circulating levels of the hormone occurs at the onset of parturition, suggesting that, in addition to its metabolic functions, this protein may act as a trigger for parturition."[24]
GeneID: 1843 DUSP1 dual specificity phosphatase 1. Also known as HVH1; MKP1; CL100; MKP-1; PTPN10. "The protein encoded by this gene is a phosphatase with dual specificity for tyrosine and threonine. The encoded protein can dephosphorylate MAP kinase MAPK1/ERK2, which results in its involvement in several cellular processes. This protein appears to play an important role in the human cellular response to environmental stress as well as in the negative regulation of cellular proliferation. Finally, the encoded protein can make some solid tumors resistant to both chemotherapy and radiotherapy, making it a target for cancer therapy."[25]
Although "macrophage proliferation and activation induce MKP-1 with different kinetics, gene expression is mediated by the proximal promoter sequences localized between -380 and -180bp. Mutagenesis experiments of the proximal element determined that CRE/AP-1 is required for LPS- or M-CSF-induced activation of the MKP-1 gene. Moreover, the results from gel shift analysis and chromatin immunoprecipitation indicated that c-Jun and CREB bind to the CRE/AP-1 box."[26]
The "same region, which contains a CREB/AP-1 box, is required for M-CSF- or LPS-dependent stimulation, although this stimulation is induced at different times after stimulation. [...] CREB and c-Jun are responsible for MKP-1 induction. The induction of c-Jun is correlated with the kinetics of MKP-1 induction by M-CSF or LPS."[26]
"MKP-1 expression induced by M-CSF or LPS is regulated by a CREB/AP-1 box."[26]
The "CRE/AP-1 box (TGACGTCT), which has been reported to be involved in the regulation of several genes [43], was critical."[26]
"When we mutated either the CRE or the AP-1 box, M-CSF- or LPS- dependent inducibility was lost [...]. Surprisingly, although the kinetics of inductions differed, the same region controlled M-CSF- and LPS-stimulated MKP-1 expression."[26]
"The DNA fragment spanned the promoter region from -193 to -169 bp containing the CRE/AP-1 box."[26]
"AP-1 is not a single protein but a series of dimeric basic region-leucine zipper proteins that belong to the Jun (c-Jun, JunB, JunD), Fos (c-Fos, FosB, Fra-1 and Fra2), and ATF (ATF2, LRF1/ATF3, B-ATF, JDP1, JDP2) subfamilies, which recognize either 12-otetradecanoylphorbol-13-acetate response elements (5'-TGAG/CTCA-3') or cAMP response elements (CRE, 5'-TGACGTCA-3') [44]."[26]
GeneID: 2353 FOS Fos proto-oncogene, AP-1 transcription factor subunit. Also known as p55; AP-1; C-FOS. "The Fos gene family consists of 4 members: FOS, FOSB, FOSL1, and FOSL2. These genes encode leucine zipper proteins that can dimerize with proteins of the JUN family, thereby forming the transcription factor complex AP-1. As such, the FOS proteins have been implicated as regulators of cell proliferation, differentiation, and transformation. In some cases, expression of the FOS gene has also been associated with apoptotic cell death."[27]
GeneID: 5179 PENK proenkephalin. Also known as PE; PENK-A. "This gene encodes a preproprotein that is proteolytically processed to generate multiple protein products. These products include the pentapeptide opioids Met-enkephalin and Leu-enkephalin, which are stored in synaptic vesicles, then released into the synapse where they bind to mu- and delta-opioid receptors to modulate the perception of pain. Other non-opioid cleavage products may function in distinct biological activities."[28]
Two forms of enkephalin have been found, one containing leucine, and the other containing methionine, both are products of the proenkephalin gene.[29]
The met-enkephalin peptide sequence is coded for by the enkephalin gene; the leu-enkephalin peptide sequence is coded for by both the enkephalin gene and the dynorphin gene.[30]
During a stress response, several Met-enkephalin analogs had increased activity in the hippocampus, while Leu-enkephalin analogs as well as somatostatins were downregulated during stress; hence, stressors impact neuropeptides, and their action is localized to a specific brain region.[31]
GeneID: 5187 PER1 period circadian regulator 1. Also known as PER; hPER; RIGUI. "This gene is a member of the Period family of genes and is expressed in a circadian pattern in the suprachiasmatic nucleus, the primary circadian pacemaker in the mammalian brain. Genes in this family encode components of the circadian rhythms of locomotor activity, metabolism, and behavior. This gene is upregulated by CLOCK/ARNTL heterodimers but then represses this upregulation in a feedback loop using PER/CRY heterodimers to interact with CLOCK/ARNTL. Polymorphisms in this gene may increase the risk of getting certain cancers. Alternative splicing has been observed in this gene; however, these variants have not been fully described."[32]
GeneID: 6750 SST somatostatin. Also known as SMST. "The hormone somatostatin has active 14 aa and 28 aa forms that are produced by alternate cleavage of the single preproprotein encoded by this gene. Somatostatin is expressed throughout the body and inhibits the release of numerous secondary hormones by binding to high-affinity G-protein-coupled somatostatin receptors. This hormone is an important regulator of the endocrine system through its interactions with pituitary growth hormone, thyroid stimulating hormone, and most hormones of the gastrointestinal tract. Somatostatin also affects rates of neurotransmission in the central nervous system and proliferation of both normal and tumorigenic cells."[33]
GeneID: 7054 TH tyrosine hydroxylase. Also known as TYH; DYT14; DYT5b. "The protein encoded by this gene is involved in the conversion of tyrosine to dopamine. It is the rate-limiting enzyme in the synthesis of catecholamines, hence plays a key role in the physiology of adrenergic neurons. Mutations in this gene have been associated with autosomal recessive Segawa syndrome. Alternatively spliced transcript variants encoding different isoforms have been noted for this gene."[34]
GeneID: 7425 VGF VGF nerve growth factor inducible. Also known as SCG7; SgVII. "This gene is specifically expressed in a subpopulation of neuroendocrine cells, and is upregulated by nerve growth factor. The structural organization of this gene is similar to that of the rat gene, and both the translated and the untranslated regions show a high degree of sequence similarity to the rat gene. The encoded secretory protein also shares similarities with the secretogranin/chromogranin family, however, its exact function is not known."[35]
GeneID: 8864 PER2 period circadian regulator 2. Also known as FASPS; FASPS1. "This gene is a member of the Period family of genes and is expressed in a circadian pattern in the suprachiasmatic nucleus, the primary circadian pacemaker in the mammalian brain. Genes in this family encode components of the circadian rhythms of locomotor activity, metabolism, and behavior. This gene is upregulated by CLOCK/ARNTL heterodimers but then represses this upregulation in a feedback loop using PER/CRY heterodimers to interact with CLOCK/ARNTL. Polymorphisms in this gene may increase the risk of getting certain cancers and have been linked to sleep disorders."[36]
Laboratory reports
Below is an outline for sections of a report, paper, manuscript, log book entry, or lab book entry. You may create your own, of course.
CRE box transcription laboratory
by --Marshallsumter (discuss • contribs) 22:00, 1 October 2018 (UTC)
Abstract
A computer experiment has been conducted on the nucleotides between the human genes ZSCAN22 and A1BG and ZNF497 and A1BG using the nucleotide sequences of GeneID:1 in the NCBI database. Using the known consensus sequence for the CRE box, one CRE box on the negative strand pointing toward A1BG in the proximal promoter in the negative direction between A1BG and ZSCAN22 was found that could be involved in the transcription of A1BG probably with an Inr rather than a TATA box. This tentatively proves that A1BG can be transcribed by a CRE box.
Introduction
According to one source, A1BG is transcribed from the direction of ZNF497: 3' - 58864890: CGAGCCACCCCACCGCCCTCCCTTGG+1GGCCTCATTGCTGCAGACGCTCACCCCAGACACTCACTGCACCGGAGTGAGCGCGACCATCATG : 58866601-5', per Michael David Winther, Leah Christine Knickle, Martin Haardt, Stephen John Allen, Andre Ponton, Roberto Justo De Antueno, Kenneth Jenkins, Solomon O. Nwaka, and Y. Paul Goldberg, Fat Regulated Genes, Uses Thereof and Compounds for Mudulating Same, US Patent Office, July 29, 2004, at http://www.google.com/patents?hl=en&lr=&vid=USPATAPP10416914&id=7iaVAAAAEBAJ&oi=fnd&printsec=abstract#v=onepage&q&f=false where the second 'G' at left of four Gs in a row is the TSS. Transcription was triggered in cell cultures and the transcription start site was found using reverse transcriptase. But, the mechanism for transcription is unknown.
Controlling the transcription of A1BG may have significant immune function against snake envenomation. A1BG forms a complex that is similar to those formed between toxins from snake venom and A1BG-like plasma proteins. These inhibit the toxic effect of snake venom metalloproteinases or myotoxins and protect the animal from envenomation.[37]
Transcription factors
Many transcription factors (TFs) may occur upstream and occasionally downstream of the transcription start site (TSS), in this gene's promoter. The following have been examined so far: (1) AGC boxes (GCC boxes), (2) ATA boxes, (3) CAAT boxes, (4) C and D boxes, (5) CArG boxes, (6) CENP-B boxes, (7) CGCG boxes, (8) DREB boxes, (9) EIF4E basal elements (4EBEs), (10) enhancer boxes (E boxes), (11) Factor II B recognition elements, (12) G boxes, (13) GLM boxes, (14) HNF6s, (15) HY boxes, (16) Metal responsive elements (MREs), (17) Motif ten elements (MTEs), (18) STAT5s, (19) TATA boxes, (20) TATCCAC boxes, (21) X boxes and (22) Y boxes.
AGC boxes (GCC boxes)
An AGC box was found in the distal promoter of either gene ZSCAN22 or A1BG on both the template and coding strands. But, as the only known transcription of A1BG occurs between Gene ID: 162968 ZNF497 and Gene ID: 1 A1BG, it is unlikely that this AGC box is naturally used to transcribe A1BG.
A full web search produced several references including a GeneCard[38] for "zinc finger protein 497" and "GCC box", including "May be involved in transcriptional regulation."[38] Zinc fingers are mentioned in association with GCC boxes in plants. It seems unlikely that an AGC box is involved in any way with the transcription of A1BG.
An extension of the nucleotide data for the positive direction from ZNF475 toward A1BG from 958 nts to 4445 nts has not discovered any AGC boxes even in the distal promoter just beyond ZNF497.
ATA boxes
Regarding hypothesis 1: there are no ATA boxes in the core promoter of A1BG from either direction or strand. This hypothesis has been shown to be true. A corollary hypothesis might be 1.1: there are no ATA boxes in the proximal promoter of A1BG from either direction or strand. This corollary hypothesis may be true. "The analysis of the promoter region indicated that a putative ATA box is located 54 nucleotides upstream from the transcription start site".[39] There is one inverse and inverse complement ATA box in the proximal promoter in the positive direction between 4050 and 4300: 3'-AAATAA-5' at 4142, and 3'-TTTATT-5' at 4142. As the TSS is at 4300 nts, this ATA box is some 158 nts away, where with the smaller data set 3'-TTTATT-5' was at 703. As the TSS is at 858 nts, this ATA box is some 155 nts away, which is approximately the same number of nts from the TSS but not close enough to be in the core promoter and not 54 nts upstream from the TSS or to match other such genes with an ATA box.
But the ATA box at 2347 is likely involved in transcription of A1BG in analogy to the rat. Although this has not been confirmed as involved, the existence of this ATA box likely proves hypothesis 1 false.
Regarding hypothesis 2: ATA boxes have a role as downstream signal transducers in A1BG. There is the following inverse ATA box on the negative strand, negative direction: 3'-AAATAA-5' at 4537. On this strand, in this direction the TSS is at 4460 nts from ZSCAN22. This ATA box is 77 nts downstream. So far no published research has been found to verify this type of downstream promoter or enhancer ATA box. There may be another isoform TSS nearby. As such, hypothesis 2 may be true.
Regarding hypothesis 3: ATA boxes may assist transcription of A1BG by other transcription factors. This hypothesis has been shown by literature search to be true. But, none of the ATA boxes for A1BG are close enough to any STAT5 promoter to match known transcription initiation.
CAAT boxes
No CAAT boxes occur on either side of A1BG.
C and D boxes
Regarding hypothesis 1: The C and D boxes are not involved in the transcription of A1BG.
There are no C boxes or D boxes in the core promoter from approximately 4425 to the possible transcription start site at nucleotide number 4460.
There are no C boxes or D boxes in the core promoter from approximately 4266 to the possible transcription start site at nucleotide number 4300.
There are no C boxes or D boxes in the proximal promoter beginning about nucleotide number 4210 in the negative direction.
There is one C box 3'-ACATCA-5' at 4116 but no D boxes in the proximal promoter beginning about nucleotide number 4050 in the positive direction.
There are four C boxes in the distal promoter: 3'-AGTAGT-5' at 2888, 3'-AGTAGT-5' at 2944, 3'-AGTAGT-5' at 3418, and 3'-AGTAGT-5' at 3521 on the negative strand in the negative direction and its complement on the positive strand.
There is one D box in the distal promoter: 3'-AGTCTG-5' at 2947 on the negative strand in the negative direction and its complement on the positive strand.
There is one C box in the distal promoter: 3'-TCATCA-5' at 3251 on the negative strand in the positive direction and its complement on the positive strand.
There is one D box in the distal promoter: 3'-AGTCTG-5' at 3923 on the negative strand in the positive direction and its complement on the positive strand.
Regarding hypothesis 2: If involved they assist transcription by other TFs.
A Google scholar search using key words: "C box", "D box", and A1BG produced zero results.
Regarding hypothesis 3: C and D boxes occur only in the proximal promoter.
GeneID: 60674 GAS5 growth arrest specific 5 (non-protein coding). "This gene produces a spliced long non-coding RNA and is a member of the 5' terminal oligo-pyrimidine class of genes. It is a small nucleolar RNA host gene, containing multiple C/D box snoRNA genes in its introns. Part of the secondary RNA structure of the encoded transcript mimics glucocorticoid response element (GRE) which means it can bind to the DNA binding domain of the glucocorticoid receptor (nuclear receptor subfamily 3, group C, member 1). This action blocks the glucocorticoid receptor from being activated and thereby stops it from regulating the transcription of its target genes. This transcript is also thought to regulate the transcriptional activity of other receptors, such as androgen, progesterone and mineralocorticoid receptors, that can bind to its GRE mimic region. Multiple functions have been associated with this transcript, including cellular growth arrest and apoptosis. It has also been identified as a potential tumor suppressor, with its down-regulation associated with cancer in multiple different tissues."[40]
"The antisense elements located immediately upstream of the D box and/or the D′ box match the sequence of the target RNA, while the areas immediately upstream of the C box and immediately downstream of the D box form a 5′–3′ terminal stem".[41]
"Small nucleolar RNAs (snoRNAs) are noncoding RNAs involved in the processing and modification of ribosomal RNAs. They are grouped in two distinct families, the box C/D family, which catalyzes methylation of 2′-hydroxyls of the pre-rRNA precursor, and the box H/ACA family, which catalyzes the modification of uridines into pseudouridines in various RNAs (reviewed in Refs. [24] and [40])."[42]
"Small nucleolar RNAs (snoRNAs) are 60–300-nucleotide-long RNAs located in the nucleolus or in Cajal bodies. They constitute one of the most abundant classes of ncRNAs [9]. Predominantly intronic, 300 different snoRNA sequences are located in the human genome. They are classified into two categories, those containing boxes C and D; and, those containing boxes H and ACA. snoRNAs are generated after splicing, debranching, and trimming of mRNA introns. Subsequently, mature snoRNAs associate with proteins to form small nucleolar ribonucleoproteins (snoRNPs). These complexes are exported into the nucleolus to participate in rRNA processing [5]."[43]
Tiny "RNAs with a modal length of 18 nt [...] map within -60 to +120 nt of transcription start sites (TSSs) in human, chicken and Drosophila. These transcription initiation RNAs (tiRNAs) are derived from sequences on the same strand as the TSS and are preferentially associated with G+C-rich promoters. The 5' ends of tiRNAs show peak density 10-30 nt downstream of TSSs, indicating that they are processed. tiRNAs are generally, although not exclusively, associated with highly expressed transcripts and sites of RNA polymerase II binding."[44]
"With exception of U3 all box C/D snoRNAs presented in this study are intron-encoded, as it is the general pathway for the biogenesis of this class of snoRNAs (22)."[45]
"Box C/D snoRNAs [...] contain conserved Box C (UGAUGA) and Box D (CUGA) elements located closely to the 5′- and 3′-ends, respectively. Internal copies of these elements are termed Box C′ and Box D′ (20,21)."[45]
Gene ID: 7422 VEGFA vascular endothelial growth factor A. "This gene is a member of the PDGF/VEGF growth factor family. It encodes a heparin-binding protein, which exists as a disulfide-linked homodimer. This growth factor induces proliferation and migration of vascular endothelial cells, and is essential for both physiological and pathological angiogenesis. Disruption of this gene in mice resulted in abnormal embryonic blood vessel formation. This gene is upregulated in many known tumors and its expression is correlated with tumor stage and progression. Elevated levels of this protein are found in patients with POEMS syndrome, also known as Crow-Fukase syndrome. Allelic variants of this gene have been associated with microvascular complications of diabetes 1 (MVCD1) and atherosclerosis. Alternatively spliced transcript variants encoding different isoforms have been described. There is also evidence for alternative translation initiation from upstream non-AUG (CUG) codons resulting in additional isoforms. A recent study showed that a C-terminally extended isoform is produced by use of an alternative in-frame translation termination codon via a stop codon readthrough mechanism, and that this isoform is antiangiogenic. Expression of some isoforms derived from the AUG start codon is regulated by a small upstream open reading frame, which is located within an internal ribosome entry site."[46]
CArG boxes
By combining a literature search with computer analysis of each promoter between ZSCAN22 and A1BG and ZNF497 and A1BG, CArG boxes have been found. To show that these CArG boxes may be used during or for transcription of A1BG at least one transcription factor has been affirmed.
A literature search of more recent results discovered: "Of the [Flowering Locus C] FLC binding sites, 69% contained at least one CArG-box motif with the core consensus sequence CCAAAAAT(G/A)G and an AAA extension at the 3′ end [. Three] other MADS-box flowering-time regulators, SOC1, SVP, and AGAMOUS-LIKE 24 (AGL24), bind to two different CArG-box motifs at 502 bp (CTAAATATGG) and 287 bp (CAATAATTGG) upstream of the translation start in the SEP3 gene (24), consistent with different specificities for the different MADS-box proteins."[47]
These together with the core motif CC(A/T)6GG suggest a more general CArG-box motif of (C(C/A/T)(A/T)6(A/G)G). Subsequent computer-program testing revealed two more general CArG boxes: 3'-CAAAAAAAAG-5' at 1399 nts from ZSCAN22 and 3'-CATTAAAAGG-5' at 3441 nts from ZSCAN22, but none within 4300 nts toward A1BG from ZNF497.
These results show that the presence of CArG boxes on the ZSCAN22 side of A1BG implies their use when transcribing A1BG, although they may be pointing toward ZSCAN22. These suggest that the hypothesis (A1BG is not transcribed by a CArG box) is false. Regarding the second hypothesis (The lack of a CArG box on either side of A1BG does not prove that it is not actively used to transcribe A1BG), the presence of more general CArG boxes in the distal promoter tentatively confirms this hypothesis.
CArG boxes do occur in the distal promoter of A1BG on the ZSCAN22 side only. And, it is likely that a CArG box is involved in some way with the transcription of A1BG.
CENP-B boxes
No CENP-B boxes occur on either side of A1BG.
CGCG boxes
On the negative strand in the negative direction (from ZSCAN22 to A1BG), looking for 3'-(A/C/G)CGCG(C/G/T)-5', there no CGCG boxes in the core promoter.
On the negative strand in the positive direction (from ZNF497 to A1BG), looking for 3'-(A/C/G)CGCG(C/G/T)-5', there no CGCG boxes in the core promoter.
There are no CGCG boxes in the negative direction of the proximal promoter.
There are no CGCG boxes in the positive direction of the other proximal promoter.
There are no CGCG boxes after nucleotide number 2460 in the negative direction of the distal promoter.
There are no CGCG boxes after nucleotide number 2300 in the positive direction of the other distal promoter.
All of the CGCG boxes found are more closely associated with ZSCAN22 or ZNF497 than A1BG.
DREB boxes
No DREB boxes occur on either side of A1BG.
EIF4E basal elements
No EIF4E basal elements, also eIF4E or (4EBE), occur on either side of A1BG.
Enhancer boxes
The presence of many enhancer boxes on both sides of A1BG demonstrate that the hypothesis: "A1BG is not transcribed by an enhancer box", is false.
The finding by literature search of evidence verifying that at least one transcription factor can enhance or inhibit the transcription of A1BG using one or more enhancer boxes disproves the hypothesis: "Existence of an enhancer box on either side of A1BG does not prove that it is actively used to transcribe A1BG".
Enhancer boxes do occur in the proximal and distal promoters of A1BG. And, it is likely that an enhancer box is involved in some way with the transcription of A1BG.
Factor II B recognition elements
Regarding hypothesis 1: B recognition element (BREu) is not involved in the transcription of A1BG.
In the negative direction, there are no BREs (BREu) in the core promoter from approximately 4425 to the possible transcription start site at nucleotide number 4460.
In the positive direction, there are no BREs (BREu) in the core promoter from approximately 4266 to the possible transcription start site at nucleotide number 4300.
There are no BREs (BREu) in the proximal promoter beginning about nucleotide number 4210 in the negative direction.
There are no BREs (BREu) in the proximal promoter beginning about nucleotide number 4050 in the positive direction.
There is one BREu in the distal promoter: 3'-CCGCACC-5' at 3047 on the negative strand in the negative direction and its complement on the positive strand.
There is one BRE in the distal promoter: 3'-CCGCACC-5' at 2566 on the negative strand in the positive direction and its complement on the positive strand.
Regarding hypothesis 2: If involved it assists transcription by other TFs.
A search of Google Scholar and the full web failed to produce any examples of BREu assisted A1BG transcription.
"A computational study based on statistical analysis of curated promoter sets concluded that up to 25% of human core promoters contain a potential BREu. The motif was found to be enriched in CpG promoters (>30% frequency) but depleted in CpG-less promoters (<10% frequency) [14]."[48]
G boxes
No G boxes occur on either side of A1BG.
GLM boxes
No GLM boxes occur on either side of A1BG.
HNF6s
HNF6s may have a downstream proximal promoter element if the computer nts sampling is additionally, approximately at least 250 nts downstream of the transcription start site. "Downstream" can refer to downstream from an enhancer but before the transcription start site, downstream from a TATA box or an initiator element but before the transcription start site (TSS), downstream from another promoter element and containing the TSS, or downstream after the TSS. The computer programs written to test for HNF6 promoters were limited to 100 nts below the apparent TSSs.
There is a HNF6 on the negative strand in the positive direction (from ZNF497 to A1BG) of 3'-TTCCGGGAA-5' at 808 in the proximal promoter, where the TSS is at 858 nts from ZNF497.
There is no such "downstream" promoter between ZSCAN22 and A1BG.
Both a TATA box or an Inr are within the core promoter. There are no HNF6s within any core promoters per the computer program sampling from ZNF497 or ZSCAN22 and A1BG.
There are no HNF6s within any core promoters per the computer program sampling from ZNF497 or ZSCAN22 and A1BG containing either TSS.
No HNF6s were detected at least to 100 nts downstream of each TSS.
There is a HNF6 on the negative strand in the positive direction (from ZNF497 to A1BG) of 3'-TTCCGGGAA-5' at 808 in the proximal promoter, where the TSS is at 858 nts from ZNF497. This direction is the only confirmed transcription of A1BG; therefore, it is likely A1BG is transcribed using this HNF6 transcription factor.
There are two HNF6s on the negative strand in the negative direction, 3'-AAGCAACTT-5' at 3506 and 3'-AAGGGACTT-5' at 3782. Both of these are in the distal promoter between ZSCAN22 and A1BG.
The only known TSS for A1BG lies at 4300 nts from just beyond ZNF497 toward A1BG. There two HNF6s in the proximal promoter between 4050 and 4300, 3'-TTATTGATTA-5' at 4164 and 3'-TATAATTGTT-5' at 4172, i.e. outside from 4242 (-58) to 4250 (-50). This suggests that HNF6 assists in the transcription of A1BG, but not downstream of the TSS.
{{fairuse}}Both "the 2.3 kb and the 160 bp proximal parts of the a1bg promoter direct sex-specific expression of the reporter gene, and that a negative regulatory element resides in the −1 kb to −160 bp region."[49]
"Computer analysis of the 2.3 kb rat a1bg promoter fragment revealed two putative HNF6 sites and one [hepatic nuclear factor 6] HNF6/HNF3 binding site at −2077/−2069, −69/−61 and −137/−128 respectively [...]."[49]
The "GH-dependent sexually dimorphic expression conveyed by the 2.3 kb a1bg promoter is enhanced by the HNF6/HNF3 site [...]."[49]
"HNF6 bound to the a1bg HNF6 oligonucleotide, but in this case, the mutated oligonucleotide was able to compete for binding when added in large excess [...]. However, [...] the HNF6 binding capacity of the mutated oligonucleotide was clearly reduced. A 20 molar excess of the mutated oligonucleotide had only a marginal effect on the binding of HNF6 [...], whereas a 20 molar excess of unlabelled probe [...] completely abolished binding. Supershift analysis with an HNF6 antibody revealed a complex with a slightly lower mobility than the HNF6 complex [...]. By extending the electrophoresis run and including nuclear extract from hypophysectomized rats, devoid of GH and thereby lacking HNF6 (Lahuna et al. 1997), the two different complexes were clearly visualized. The complex with the lower mobility is most probably due to the binding of HNF3, in analogy with what was shown by Lahuna et al. for the CYP2C12 HNF6 binding site; HNF3 can bind to the site in the absence of HNF6 (Lahuna et al. 1997). [...] HNF6 could bind to their respective site in the a1bg promoter in vitro, and the mutations introduced in respective site abolished binding of the corresponding factor."[49]
The "expression of a −116/−89 deletion construct in which also the HNF6 site was mutated, (−116/−89) delmutHNF6-Luc, [...] the generated luciferase activities were reduced in both sexes [...]. This is in contrast to that mutation/deletion of the sites separately only affected the expression in female livers."[49]
The "−116/−89 region contains a site(s) of importance for the GH-dependent and female-specific expression of the a1bg gene, and that the impact of this region together with the HNF6 site is more complex than mere enhancement of the expression in females."[49]
Following "mutation of the HNF6-binding element, mutHNF6-Luc, the sex-differentiated expression was attenuated due to reduced expression in females. Thus, for a1bg, the sex-related difference in amount of HNF6 is likely to contribute to the sex-differentiated and female characteristic expression."[49]
Nuclear "proteins binding to the a1bg −116/−89 region [are] members of the [nuclear factor 1] NF1 and the [octamer transcription factor] Oct families of transcription factors. NF1 genes are expressed in most adult tissues (Osada et al. 1999). It is not known how NF1 modulates transcriptional activity, and both activation and repression of transcription have been reported (Gronostajski 2000). Cofactors such as CBP/p300 and HDAC have been shown to interact with NF1 proteins suggesting modulation of chromatin structure (Chaudhry et al. 1999). NF1 factors have also been shown to interact directly with the basal transcription machinery as well as with other transcription factors, including Stat5 (Kim & Roeder 1994, Mukhopadhyay et al. 2001) and synergistic effects with HNF4 have been reported (Ulvila et al. 2004). In addition to the HNF6, Stat5 and NF1/Oct sites, the a1bg promoter harbours an imperfect HNF4 site at −51/−39 with two mismatches compared with the HNF4 consensus site. HNF4 is clearly important for the expression of CYP2C12 (Sasaki et al. 1999), however, the −51/−39 region in a1bg was not protected in the footprinting analysis and was therefore not analysed further. Like NF1, Oct proteins have been reported to be involved in activation as well as repression of gene expression (Phillips & Luisi 2000). [...] Moreover, NF1 and Oct-1 have been shown to, reciprocally, facilitate each other’s binding (O’Connor & Bernard 1995, Belikov et al. 2004)."[49]
In the diagram on the right is liver "expression of a1bg-luciferase constructs. (A) Stat5 and HNF6 consensus sequences and corresponding sites in the 2.3 kb a1bg promoter alongside with the used mutations. (B) Female (black bars) and male (open bars) rats [results]."[49]
"Computer analysis of the 2.3 kb rat a1bg promoter fragment revealed [a] HNF6 [site] at [...] −69/−61 [...]."[49]
The murine downstream promoter element is only 11 nts displaced from the human one. This suggests a HNF6 participation in human gene transcription of A1BG.
"Computer analysis of the 2.3 kb rat a1bg promoter fragment revealed two putative HNF6 sites [...] at −2077/−2069 [and] −69/−61 [...]."[49]
There are two HNF6s on the negative strand in the negative direction, 3'-AAGCAACTT-5' at 3506 (-954) and 3'-AAGGGACTT-5' at 3782 (-678) in the distal promoter between ZSCAN22 and A1BG. Although much closer than their likely murine counterparts, they are on the other side of A1BG from the HNF6 site confirming hypothesis 1. If active in humans or murine-like HNF6s occur within or beyond ZNF497 in this distal promoter, then human A1BG is transcribed using HNF6 promoters disproving hypothesis 2.
A Google Scholar search using ZNF497 with HNF6 found no articles discussing HNF6 sites inside or associated with ZNF497. To confirm they exist, a data file going 4300 nts to just beyond ZNF497 has been created and tested for a distal promoter on this side. Distal HNF6s in the positive direction, if they exist, would be inside ZNF497 or beyond, e.g., 3'-ATGTCCATGG-5' at 3581 was found.
Literature search has found that HNF6s assist transcription of A1BG by other transcription factors.[49] The proximal HNF6 promoter is -58 to -50 from A1BG TSS. If another HNF6 promoter is at -2.3 kb, it is about -1.4 kb inside ZNF497 which is 3212 nts long. Per analogy to the rat this would be expected.[49]
Per earlier laboratories transcription factors may occur in the distal promoters on the ZNF497 side of A1BG for
- ATA boxes 3'-AATAAA-5' occurs at 3427,
- CArG boxes,
- Enhancer boxes,
- HY boxes,
- MREs and
- STAT5s 3'-TTCCATGAA-5' occurs at 128.
The HNF6 promoter on the other side of A1BG (at about +3 kb is way beyond -2.1 through ZNF497 unless the DNA is folded to allow the HNF6 on the ZSCAN22 side to be used in analogy to the HNF6 on the same side as in the rat.[49]
HNF6s have a role as downstream signal transducers in A1BG, where the murine downstream promoter element is only 11 nts displaced from the human one. This suggests a HNF6 participation in human gene transcription of A1BG.
HY boxes
HY boxes were not found in either core promoters or the proximal promoters in either direction. However, HY boxes were found in the distal promoters on both sides of A1BG. No genes are described in the literature so far as transcribed from HY boxes in any distal promoters.
Either A1BG can be transcribed by HY boxes in the distal promoter, or A1BG is not transcribed by HY boxes. As the literature appears absent from a Google Scholar advanced search to confirm possible transcription from distal promoters, wet chemistry experiments are needed to test the possibility.
Metal responsive elements (MREs)
By combining a literature search with computer analysis of the promoter between ZSCAN22 and A1BG and ZNF497 and A1BG, metal responsive elements have been found. Literature search has also discovered at least three post-translational isoforms including the unaltered precursor. Although no metal responsive elements overlap any enhancer boxes in the distal promoter, there are elements in the distal promoter.
"The human genome is estimated to contain 700 zinc-finger genes, which perform many key functions, including regulating transcription. [Four] clusters of zinc-finger genes [occur] on human chromosome 19".[50]
Nearby zinc-fingers on chromosome 19 include ZNF497 (GeneID: 162968), ZNF837 (GeneID: 116412), and ZNF8 (GeneID: 7554).
"In rodents and in humans, about one third of the zinc-finger genes carry the Krüppel-associated box (KRAB), a potent repressor of transcription (Margolin et al. 1994), [...]. There are more than 200 KRAB-containing zinc-finger genes in the human genome, about 40% of which reside on chromosome 19 and show a clustered organization suggesting an evolutionary history of duplication events (Dehal et al. 2001)."[50]
ZNF8 is in cluster V along with A1BG.[50]
"In contrast to the four clusters considered [I through IV], one that occurs at the telomere of chromosome 19, which we will call cluster V, has been very stable [over mouse, rat, and human]."[50]
"Apart from the somewhat unexpected location of Zfp35 on mouse chromosome 18 and of the AIBG orthologs on mouse chromosome 15 and rat chromosome 7, there has been little rearrangement."[50]
So far no article has reported any linkage between zinc, including various zinc fingers, or cadmium, and A1BG.
Regarding additional isoforms, mention has been made of "new genetic variants of A1BG."[51]
"Proteomic analysis revealed that [a circulating] set of plasma proteins was α 1 B-glycoprotein (A1BG) and its post-translationally modified isoforms."[52]
Pharmacogenomic variants have been reported. There are A1BG genotypes.[53]
A1BG has a genetic risk score of rs893184.[53]
"A genetic risk score, including rs16982743, rs893184, and rs4525 in F5, was significantly associated with treatment-related adverse cardiovascular outcomes in whites and Hispanics from the INVEST study and in the Nordic Diltiazem study (meta-analysis interaction P=2.39×10−5)."[53]
"rs893184 causes a histidine (His) to arginine (Arg) [nonsynonymous single nucleotide polymorphism (nsSNP), A (minor) for G (major)] substitution at amino acid position 52 in A1BG."[53]
For example, GeneID: 9 has isoforms: a, b, X1, and X2. Each of these (a and b) have variants. Variants 1-6 and 9 all encode the same isoform (a).
Variants 7, 8 and 10 all encode isoform b. Isoforms X1 and X2 are predicted.
Variants can differ in promoters, untranslated regions, or exons. For GeneID: 9: This variant (1) represents the longest transcript but encodes the shorter isoform (a). This variant is transcribed from a promoter known as P1, promoter 2, or NATb promoter.
This variant (2, also known as Type IID) lacks an alternate exon in the 5' UTR, compared to variant 1. This variant is transcribed from a promoter known as P1, promoter 2, or NATb promoter.
This variant (9, also known as Type IA) has a distinct 5' UTR and represents use of an alternate promoter known as the NATa or P3 promoter, compared to variant 1.
But, A1BG in NCBI Gene lists only one isoform, the gene locus itself, and the protein transcribed is a precursor subject to translational or more likely post-translational modifications.
The presence of multiple MREs coupled with experimental results from the literature indicating post-translational isoforms tends to confirm the existence of two or more isoforms for A1BG.
It isn't known which, if any, assist in locating and affixing the transcription mechanism for A1BG. This examination is the first to test one such DNA-occurring TF: the HNF6s.
The presence of multiple MREs coupled with experimental results from the literature indicating post-translational isoforms tends to confirm the existence of two or more isoforms for A1BG and likely transcription from either side.
Motif ten elements
No Motif ten elements occur on either side of A1BG.
STAT5s
STAT5s have a role as downstream signal transducers in A1BG, where the murine downstream promoter element is only 11 nts displaced from the human one. This suggests a STAT5 participation in human gene transcription of A1BG in the proximal promoter downstream between any other promoter and the TSS on the ZNF497 side of A1BG.
A1BG is not transcribed by any STAT5s is clearly disproved by the STAT5 transcription factor in the proximal promoter on the ZNF497 side of A1BG.
STAT5s may assist transcription of A1BG by other transcription factors, literature search has found that STAT5s assist transcription of A1BG by other transcription factors.[49] The proximal STAT5 promoter is -58 to -50 from A1BG TSS. If another STAT5 promoter is at -2.3 kb, it is about -1.4 kb inside ZNF497 which is 3212 nts long. Per analogy to the rat this would be expected.[49] A STAT5 transcription site lies at 3'-TTCCGGGAA-5' at 4247 in the proximal promoter, i.e. from 4242 (-58) to 4250 (-50). This suggests that STAT5 assists in the transcription of A1BG.
TATA boxes
On the negative strand in the negative direction (from ZSCAN22 to A1BG), looking for 3'-TATA-A/T-A-A/T-A/G-5', there no TATA boxes in the core promoter.
On the negative strand in the positive direction (from ZNF497 to A1BG), looking for 3'-TATA-A/T-A-A/T-A/G-5-5', there no TATA boxes in the core promoter.
There are no TATA boxes in the negative direction of the proximal promoter.
There are no TATA boxes in the positive direction of the proximal promoter.
For the positive strand in the negative direction looking for 3'-TATA-A/T-A-A/T-A/G-5', there's one 3'-TATATAAA-5' at 2874 nts, its complement and inverse complement of the distal promoter.
Any TATA boxes before A1BG from the direction of ZN497 may be out to 2300 nts. None were found in the distal promoter.
On the positive strand, in the nucleotide region between gene ZSCAN22 (NCBI GeneID: 342945) and A1BG (NCBI GeneID: 1) are 211 TATA box-like 8 nt long sequences. Of these,
- TATAAAAG occurs at 58853713 + 183 nts and
- TATAAAAG at 58853713 + 222. This is a TATA box found with some genes.[54] But, the optimal TBP recognition sequence 3'-TATATAAG-5',[55] does not occur.
- TATATAAA occurs only once at 2874 nts from the end of ZSCAN22. TBP is bound to this sequence and TATAAAAG above.[56][57]
- TATAAA occurs seven times, with the closest one at 2874 nts from the end of ZSCAN22. "In virtually every RNA polymerase II-transcribed gene examined, the sequence TATAAA was present 25 to 30 nts upstream of the transcription start site."[12]
A1BG does not have a TATA box in the core promoter region. There is the sequence 3'-TGCTATATAGATGGCAACTAAGCACTTGGGGAAAAAA-5' for which the first nt (T) is number 58856598 or 1574 nt upstream from the beginning of the 3'-UTR at 58858172. Unless another variant exists, -1574 nt from the beginning of the 3'-UTR is a large number of nts away from the TSS.
The closest TATA box-like sequence is 3'-CTCTTAAG-5' on the template strand at 4408 nts from the end of ZSCAN22, which is upstream from the core promoter.
The extra TATA boxes between ZSCAN22 and A1BG strongly suggest that there is at least one gene (or pseudogene) between ZSCAN22 and A1BG not currently in the NCBI database.
On the negative strand between ZNF497 and A1BG, there are no TATA boxes of the form 3’-TATA-A/T-A-A/T-A/G-5’.
For the negative strand going from ZSCAN22 to A1BG there are two TATA boxes: 3'-TATATATA-5' at 1600 nts and 3'-TATATAAA-5' at 1602 nts. These are way too far from the possible TSS in this direction.
These two TATA boxs in the distal promoter at approximately -2860 nts from the TSS suggest that there may be a short gene between ZSCAN22 and A1BG.
The hypothesis: TATA boxes are not involved in the transcription of A1BG is true. "In virtually every RNA polymerase II-transcribed gene examined, the sequence TATAAA was present 25 to 30 nts upstream of the transcription start site."[12]
There are no TATA boxes at all between ZNF497 and A1BG.
On the negative strand between ZSCAN22 and A1BG there are many TATA boxes between 184 nts from ZSCAN22 and 2874 nts from ZSCAN22 yet no genes are apparently known to occur between ZSCAN22 and A1BG. ZSCAN22 has several isoforms but all end exactly at the one TSS on the A1BG side.
From the number and variety of TFs on both sides of A1BG, multiple transcriptions should be possible. Any connection between bone, muscle and brain mass loss and A1BG likely uses one or more of the sides, directions, or forms (16 ways) and includes one or more TFs. Determining which produces deleterious effects is the first step toward reversal in a zero-g radiation inducing environment.
TATCCAC boxes
No TATCCAC boxes occur on either side of A1BG.
X boxes
No X boxes occur on either side of A1BG.
Y boxes
No Y boxes occur on either side of A1BG.
Experiments
The Basic programs (starting with SuccessablesCRE.bas) were written to compare nucleotide sequences with the sequences on either the template strand (-), or coding strand (+), of the DNA, in the negative direction (-), or the positive direction (+), including extending the number of nts from 958 to 4445, looking for CRE boxes, their possible complements and inverses to test the hypothesis that CRE boxes are not involved in the transcription of A1BG.
Results
On the negative strand in the negative direction (from ZSCAN22 to A1BG), looking for 3'-TGACGTCA-5', there no CRE boxes in the core promoter.
On the negative strand in the positive direction (from ZNF497 to A1BG), looking for 3'-TGACGTCA-5', there no CRE boxes in the core promoter.
There is one CRE box 3'-TGACGTCA-5' in the negative direction in the proximal promoter at 4317 nts from ZSCAN22.
There are no CRE boxes in the positive direction in the proximal promoter.
There are no CRE boxes after nucleotide number 2460 in the negative direction in the distal promoter.
There are no CRE boxes after nucleotide number 2300 in the positive direction in the distal promoter.
Discussions
One CRE box occurred from -143 to -135 nts (3'-TGACGTCA-5') upstream of the TSS on the negative strand in the negative direction between A1BG and ZSCAN22.
The "proximal positive response region [...] contains a CRE homology ((-71 bp)-TGACGTAA-(-64 bp))".[21]
"Each species' promoter contains a cAMP/response element (CRE)1 consensus sequence ([7]) upstream of a TATA box."[8]
The "isolated mouse chromogranin B promoter [specifically] the proximal chromogranin B promoter (from −216 to −91 bp); [...] contains [...] a cAMP response element (CRE; at [−102 bp]TGACGTCA[−95 bp])."[23]
"The DNA fragment spanned the promoter region from -193 to -169 bp containing the CRE/AP-1 box."[26]
While the CRE box does not overlap other CRE boxes it is in the range and pointed in the right direction to infer that A1BG can be transcribed by it even though a TATA box is not present.
A Google Scholar search using "CRE/AP-1 box" and A1BG or "CRE box" and A1BG produced no results. It appears unlikely that anyone has tried to transcribe A1BG using this CRE box.
"The human [Transforming growth factor b1] TGFB1 promoter region contains two binding sequences for [Activator protein-1] AP-1, designated AP-1 box A (TGACTCT) and box B (TGTCTCA), which mediate the upregulation of promoter activity via a PKC-dependent pathway after exposure of cells to a high-glucose environment (Refs 37, 38)."[58]
"Several transcription factors have been implicated in glucose-mediated expression of genes involved in diabetic nephropathy. This review focuses on the transcription factors upstream stimulatory factors 1 and 2 (USF1 and 2), activator protein 1 (AP-1), nuclear factor (NF)-κB, cAMP-response-element-binding protein (CREB), nuclear factor of activated T cells (NFAT), and stimulating protein 1 (Sp1). In response to high glucose, several of these transcription factors regulate the gene encoding the profibrotic cytokine transforming growth factor β, as well as genes for a range of other proteins implicated in inflammation and extracellular matrix turnover, including thrombospondin 1, the chemokine CCL2, osteopontin, fibronectin, decorin, plasminogen activator inhibitor 1 and aldose reductase."[58]
Conclusions
There is one CRE box on the negative strand pointing toward A1BG in the proximal promoter in the negative direction between A1BG and ZSCAN22 that can be involved in the transcription of A1BG probably with an Inr rather than a TATA box. This tentatively proves hypothesis 1 false; i.e., A1BG can be transcribed by a CRE box.
Laboratory evaluations
To assess your example, including your justification, analysis and discussion, I will provide such an assessment of my example for comparison and consideration.
Evaluation
No wet chemistry experiments were performed to confirm that Gene ID: 1 may be transcribed from either side using TFs in proximal promoters. The NCBI database is generalized, whereas individual human genome testing could demonstrate that A1BG is transcribed from either side TFs. Sufficient nts have been added to the data sets for the ZNF497 side to confirm likely transcription of A1BG.
See also
References
- ↑ Bourtchuladze; et al. "Deficient long-term memory in mice with a targeted mutation of the cAMP-responsive element-binding protein". Cell. 79 (1): 59–68. doi:10.1016/0092-8674(94)90400-6. PMID 7923378.
- ↑ 2.0 2.1 Purves, Dale; George J. Augustine; David Fitzpatrick; William C. Hall; Anthony-Samuel LaMantia; James O. McNamara; Leonard E. White (2008). Neuroscience (4th ed.). Sinauer Associates. pp. 170–6. ISBN 978-0-87893-697-7.
- ↑ Binding of a nuclear protein to the cyclic-AMP response element of the somatostatin gene. Montminy MR and Bilezikjian LM. Nature. 1987 Jul 9-15;328(6126):175-8.
- ↑ Dibner, Charna; Schibler, Ueli; Albrecht, Urs (2010). "The Mammalian Circadian Timing System: Organization and Coordination of Central and Peripheral Clocks". Annual Review of Physiology. 72 (1): 517–549. doi:10.1146/annurev-physiol-021909-135821. PMID 20148687. Retrieved 2015-04-09.
- ↑ Silva; et al. "CREB and Memory" (PDF). Annual Review of Neuroscience. 21: 127–148. doi:10.1146/annurev.neuro.21.1.127.
- ↑ Downregulation of CREB expression in Alzheimer's brain and in Ab-treated rat hippocampal neurons
- ↑ 7.0 7.1 7.2 7.3 7.4 Marc R. Montminy, Kevin A. Sevarino, John A. Wagner, Gail Mandel, and Richard H. Goodman (September 1986). "Identification of a cyclic-AMP-responsive element within the rat somatostatin gene" (PDF). Proceedings of the National Academy of Sciences of the USA. 83 (18): 6382–6. Retrieved 17 September 2018.
- ↑ 8.0 8.1 8.2 8.3 Hongjiang Wu, Sushil K. Mahata, Manjula Mahata, Nicholas J. G. Webster, Robert J. Parmer, and Daniel T. O'Connor (1 July 1995). "A Functional Cyclic AMP Response Element Plays a Crucial Role in Neuroendocrine Cell Type-specific Expression of the Secretory Granule Protein Chromogranin A". The Journal of Clinical Investigation. 96 (1): 568–578. doi:10.1172/JCI118069. Retrieved 17 September 2018.
- ↑ Carlezon, WA; Duman, RS; Nestler, EJ (August 2005). "The many faces of CREB". linkinghub.elsevier.com. 28: 436–445. doi:10.1016/j.tins.2005.06.005. PMID 15982754. Retrieved 2015-04-09.
- ↑ Altarejos, Judith Y.; Montminy, Marc (March 2011). "CREB and the CRTC co-activators: sensors for hormonal and metabolic signals". Nature Reviews Molecular Cell Biology. 12 (3): 141–151. doi:10.1038/nrm3072. ISSN 1471-0072. PMC 4324555. PMID 21346730. Retrieved 2015-04-09.
- ↑ Jennifer E.F. Butler, James T. Kadonaga (October 15, 2002). "The RNA polymerase II core promoter: a key component in the regulation of gene expression". Genes & Development. 16 (20): 2583–292. doi:10.1101/gad.1026202. PMID 12381658.
- ↑ 12.0 12.1 12.2 Stephen T. Smale and James T. Kadonaga (July 2003). "The RNA Polymerase II Core Promoter" (PDF). Annual Review of Biochemistry. 72 (1): 449–79. doi:10.1146/annurev.biochem.72.121801.161520. PMID 12651739. Retrieved 2012-05-07.
- ↑ Thomas Shafee and Rohan Lowe (2017). "Eukaryotic and prokaryotic gene structure" (PDF). WikiJournal of Medicine. 4 (1): 2. doi:10.15347/wjm/2017.002. Retrieved 2017-04-06. Unknown parameter
|month=ignored (help) - ↑ Koichi Takayama, Ken-ichirou Morohashi, Shin-ichlro Honda, Nobuyuki Hara and Tsuneo Omura (1994). "Contribution of Ad4BP, a Steroidogenic Cell-Specific Transcription Factor, to Regulation of the Human CYP11A and Bovine CYP11B Genes through Their Distal Promoters". The Journal of Biochemistry. 116 (1): 193–203. doi:10.1093/oxfordjournals.jbchem.a124493. Retrieved 2017-08-16. Unknown parameter
|month=ignored (help) - ↑ Michelle Craig Barton, Navid Madani, and Beverly M. Emerson (8 July 1997). "Distal enhancer regulation by promoter derepression in topologically constrained DNA in vitro". Proceedings of the National Academy of Sciences of the United States of America. 94 (14): 7257–62. Retrieved 2017-08-16.
- ↑ A Aoyama, T Tamura, K Mikoshiba (March 1990). "Regulation of brain-specific transcription of the mouse myelin basic protein gene: function of the NFI-binding site in the distal promoter". Biochemical and Biophysical Research Communications. 167 (2): 648–53. doi:10.1016/0006-291X(90)92074-A. Retrieved 2012-12-13.
- ↑ J Gao and L Tseng (June 1996). "Distal Sp3 binding sites in the hIGBP-1 gene promoter suppress transcriptional repression in decidualized human endometrial stromal cells: identification of a novel Sp3 form in decidual cells". Molecular Endocrinology. 10 (6): 613–21. doi:10.1210/me.10.6.613. Retrieved 2012-12-13.
- ↑ Peter Pasceri, Dylan Pannell, Xiumei Wu, and James Ellis (July 15, 1998). "Full activity from human β-globin locus control region transgenes requires 5′ HS1, distal β-globin promoter, and 3′ β-globin sequences". Blood. 92 (2): 653–63. Retrieved 2012-12-13.
- ↑ RefSeq (November 2015). brain derived neurotrophic factor. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 12 September 2018.
- ↑ RefSeq (September 2014). chromogranin A. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 15 September 2018.
- ↑ 21.0 21.1 21.2 21.3 21.4 21.5 21.6 21.7 Kechun Tang, Hongjiang Wu, Sushil K. Mahata, Laurent Taupenot, David J. Rozansky, Robert J. Parmer, and Daniel T. O’Connor (8 November 1996). "Stimulus-transcription Coupling in Pheochromocytoma Cells Promoter region-specific activation of chromogranin A biosynthesis" (PDF). The Journal of Biological Chemistry. 271 (45): 28382–28390. Retrieved 15 September 2018.
- ↑ RefSeq (January 2009). chromogranin B. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 12 September 2018.
- ↑ 23.0 23.1 23.2 23.3 Nitish R. Mahapatra, Manjula Mahata, Arun K. Datta, Hans-Hermann Gerdes, Wieland B. Huttner, Daniel T. O’Connor, Sushil K. Mahata (1 October 2000). "Neuroendocrine Cell Type-Specific and Inducible Expression of the Chromogranin B Gene: Crucial Role of the Proximal Promoter". Endocrinology. 141 (10): 3668–3678. doi:10.1210/endo.141.10.7725. Retrieved 15 September 2018.
- ↑ RefSeq (November 2015). corticotropin releasing hormone. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 12 September 2018.
- ↑ RefSeq (August 2017). dual specificity phosphatase 1. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 16 September 2018.
- ↑ 26.0 26.1 26.2 26.3 26.4 26.5 26.6 26.7 Cristina Casals‐Casas, Eva Álvarez, Maria Serra, Carolina de la Torre, Consol Farrera, Ester Sánchez‐Tilló, Carme Caelles, Jorge Lloberas, and Antonio Celada (7 July 2009). "CREB and AP-1 activation regulates MKP-1 induction by LPS or M-CSF and their kinetics correlate with macrophage activation versus proliferation". European Journal Immunology. 39 (7): 1902–1913. doi:10.1002/eji.200839037. Retrieved 16 September 2018.
- ↑ RefSeq (July 2008). Fos proto-oncogene, AP-1 transcription factor subunit. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 12 September 2018.
- ↑ RefSeq (12 August 2018). proenkephalin. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 12 September 2018.
- ↑ Noda M, Teranishi Y, Takahashi H, Toyosato M, Notake M, Nakanishi S, Numa S (June 1982). "Isolation and structural organization of the human preproenkephalin gene". Nature. 297 (5865): 431–4. Bibcode:1982Natur.297..431N. doi:10.1038/297431a0. PMID 6281660.
- ↑ Opioid peptides: Molecular pharmacology, biosynthesis and analysis, R.S. Rapaka and R. L. Hawks (editors) in a National Institute on Drug Abuse Research Monograph (#70), 1986.
- ↑ Henry MS, Gendron L, Tremblay ME, Drolet G (2017). "Enkephalins: Endogenous Analgesics with an Emerging Role in Stress Resilience". review. Neural Plasticity. 2017: 1546125. doi:10.1155/2017/1546125. PMC 5525068. PMID 28781901.
- ↑ RefSeq (January 2014). period circadian regulator 1. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 12 September 2018.
- ↑ RefSeq (July 2008). somatostatin. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 12 September 2018.
- ↑ RefSeq (July 2008). tyrosine hydroxylase. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 12 September 2018.
- ↑ RefSeq (12 August 2018). VGF nerve growth factor inducible. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 12 September 2018.
- ↑ RefSeq (January 2014). period circadian regulator 2. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 12 September 2018.
- ↑ Udby L, Sørensen OE, Pass J, Johnsen AH, Behrendt N, Borregaard N, Kjeldsen L. (October 2004). "Cysteine-rich secretory protein 3 is a ligand of alpha1B-glycoprotein in human plasma". Biochemistry. 43 (40): 12877–86. doi:10.1021/bi048823e. PMID 15461460. Retrieved 2011-11-28.
- ↑ 38.0 38.1 Weizmann Institute of Science (2017). Zinc Finger Protein 497. Israel: Weizmann Institute of Science. Retrieved 2017-08-20.
- ↑ Annie Charbonneau and Van-Luu The (2001). "Genomic organization of a human 5β-reductase and its pseudogene and substrate selectivity of the expressed enzyme". Biochimica et Biophysica Acta (BBA) - Gene Structure and Expression. 1517 (2): 228–235. doi:10.1016/S0167-4781(00)00278-5. Retrieved 2017-11-17. Unknown parameter
|month=ignored (help) - ↑ RefSeq (12 May 2018). GAS5 growth arrest specific 5 (non-protein coding) [ Homo sapiens (human) ]. 8600 Rockville Pike, Bethesda MD, 20894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 12 May 2018.
- ↑ Satu Nahkuri, Ryan J. Taft, Darren J. Korbie and John S. Mattick (2008). "Molecular Evolution of the HBII-52 snoRNA Cluster". Journal of Molecular Biology. 381 (4): 810–815. doi:10.1016/j.jmb.2008.06.057. Retrieved 2018-5-16. Unknown parameter
|month=ignored (help); Check date values in:|accessdate=(help) - ↑ Kevin Roy and Guillaume F. Chanfreau (2012). Eukaryotic RNases and their Partners in RNA Degradation and Biogenesis, Part A, In: The Enzymes. sciencedirect. Retrieved 16 May 2018.
- ↑ Yannick Delpu, Dorian Larrieu, Marion Gayral, Dina Arvanitis, Marlène Dufresne, Pierre Cordelier, Jérôme Torrisani (2016). Gerda Egger and Paola Arimondo, ed. Noncoding RNAs, In: Drug Discovery in Cancer Epigenetics. sciencedirect. pp. 305–326. ISBN 978-0-12-802208-5. Retrieved 16 May 2018.
- ↑ RJ Taft, EA Glazov, N Cloonan, C Simons, S Stephen, GJ Faulkner, T, Lassmann, AR Forrest, SM Grimmond, K Schroder, K Irvine, T Arakawa, M Nakamura, A Kubosaki, K Hayashida, C Kawazu, M Murata, H Nishiyori, S Fukuda, J Kawai, CO Daub, DA Hume, H Suzuki, V Orlando, P Carninci, Y Hayashizaki, JS Mattick (2009). "Tiny RNAs associated with transcription start sites in animals". Nature Genetcs. 41 (5): 572–8. doi:10.1038/ng.312. Retrieved 2018-5-16. Unknown parameter
|month=ignored (help); Check date values in:|accessdate=(help) - ↑ 45.0 45.1 Markus Brameier, Astrid Herwig, Richard Reinhardt, Lutz Walter, Jens Gruber (2011). "Human box C/D snoRNAs with miRNA like functions: expanding the range of regulatory RNAs". Nucleic Acids Research. 39 (2): 675–686. doi:10.1093/nar/gkq776. Retrieved 2018-5-16. Unknown parameter
|month=ignored (help); Check date values in:|accessdate=(help) - ↑ RefSeqNov2015 (November 2015). VEGFA vascular endothelial growth factor A [Homo sapiens (human)]. 8600 Rockville Pike, Bethesda 0894 USA: National Center for Biotechnology Information, U.S. National Library of Medicine. Retrieved 22 May 2018.
- ↑ Weiwei Deng, Hua Ying, Chris A. Helliwell, Jennifer M. Taylor, W. James Peacock, and Elizabeth S. Dennis (2011). "FLOWERING LOCUS C (FLC) regulates development pathways throughout the life cycle of Arabidopsis". Proceedings of the National Academy of Sciences United States of America. 108 (16): 6680–6685. doi:10.1073/pnas.1103175108. Retrieved 2017-09-17. Unknown parameter
|month=ignored (help) - ↑ Thomas K Albert, Korbinian Grote, Stefan Boeing and Michael Meisterernst (2010). "Basal core promoters control the equilibrium between negative cofactor 2 and preinitiation complexes in human cells". Genome Biology. 11: R33. doi:10.1186/gb-2010-11-3-r33. Unknown parameter
|month=ignored (help); Check date values in:|accessdate=(help);|access-date=requires|url=(help) - ↑ 49.00 49.01 49.02 49.03 49.04 49.05 49.06 49.07 49.08 49.09 49.10 49.11 49.12 49.13 49.14 49.15 Cissi Gardmo and Agneta Mode (2006). "In vivo transfection of rat liver discloses binding sites conveying GH-dependent and female-specific gene expression". Journal of Molecular Endocrinology. 37 (3): 433–441. doi:10.1677/jme.1.02116. Retrieved 2017-09-01. Unknown parameter
|month=ignored (help) - ↑ 50.0 50.1 50.2 50.3 50.4 Deena Schmidt and Rick Durrett (2004). "Adaptive Evolution Drives the Diversification of Zinc-Finger Binding Domains". Molecular Biology and Evolution. 21 (12): 2326–2339. doi:10.1093/molbev/msh246. Retrieved 2017-10-16. Unknown parameter
|month=ignored (help) - ↑ H Eiberg, ML Bisgaard, J Mohr (1989). "Linkage between alpha 1B-glycoprotein (A1BG) and Lutheran (LU) red blood group system: assignment to chromosome 19: new genetic variants of A1BG". Clinical genetics. 36 (6): 415–8. PMID 2591067. Retrieved 2017-10-08. Unknown parameter
|month=ignored (help) - ↑ John R. Stehle Jr., Mark E. Weeks, Kai Lin, Mark C. Willingham, Amy M. Hicks, John F. Timms, Zheng Cui (2007). "Mass spectrometry identification of circulating alpha-1-B glycoprotein, increased in aged female C57BL/6 mice". Biochimica et Biophysica Acta (BBA) - General Subjects. 1770 (1): 79–86. Retrieved 2017-10-08. Unknown parameter
|month=ignored (help) - ↑ 53.0 53.1 53.2 53.3 Caitrin W. McDonough, Yan Gong, Sandosh Padmanabhan, Ben Burkley, Taimour Y. Langaee, Olle Melander, Carl J. Pepine, Anna F. Dominiczak, Rhonda M. Cooper-DeHoff, Julie A. Johnson (2013). "Pharmacogenomic Association of Nonsynonymous SNPs in SIGLEC12, A1BG, and the Selectin Region and Cardiovascular Outcomes" (PDF). Hypertension. 62 (1): 48–54. doi:10.1161/HYPERTENSIONAHA.111.00823. PMID 23690342. Retrieved 2017-10-08. Unknown parameter
|month=ignored (help) - ↑ Georgia A. Patikoglou, Joseph L. Kim, Liping Sun, Sang-Hwa Yang, Thomas Kodadek, and Stephen K. Burley (1999). "TATA element recognition by the TATA box-binding protein has been conserved throughout evolution". Genes & Development. 13: 3217–30. Retrieved 2012-06-12.
- ↑ Jie Min Wong and Erik Bateman (25 May 1994). "TBP-DNA interactions in the minor groove discriminate between A: T and T: A base pairs". Nucleic Acids Research. 22 (10): 1890–96. doi:10.1093/nar/22.10.1890. Retrieved 2012-06-12.
- ↑ Joseph L. Kim, Dimitar B. Nikolov & Stephen K. Burley (7 October 1993). "Co-crystal structure of TBP recognizing the minor groove of a TATA element". Nature. 365: 520–7. doi:10.1038/365520a0. Retrieved 2012-06-12.
- ↑ Youngchang Kim, James. H. Geiger, Steven Hahn & Paul B. Sigler (7 October 1993). "Crystal structure of a yeast TBP/TATA-box complex". Nature. 365: 512–20. Retrieved 2012-06-12.
- ↑ 58.0 58.1 Amber Paratore Sanchez and Kumar Sharma (July 2009). "Transcription factors in the pathogenesis of diabetic nephropathy". Expert Reviews in Molecular Medicine. 11: e13. doi:10.1017/S1462399409001057. Retrieved 1 October 2018.
External links
- CS1 maint: Explicit use of et al.
- CS1 maint: Multiple names: authors list
- Pages with citations using unsupported parameters
- CS1 errors: dates
- Pages using citations with accessdate and no URL
- Pages with broken file links
- Biochemistry laboratories
- Genetics laboratories
- Resources last modified in November 2019