Yersinia pestis
|
Yersinia pestis infection Microchapters |
|
Differentiating Yersinia Pestis Infection from other Diseases |
|---|
|
Diagnosis |
|
Treatment |
|
Case Studies |
|
Yersinia pestis On the Web |
|
American Roentgen Ray Society Images of Yersinia pestis |
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [1]; Associate Editor(s)-in-Chief: Esther Lee, M.A.; Rim Halaby, M.D. [2]; João André Alves Silva, M.D. [3]
Overview
Yersinia pestis (Y. pestis), a rod-shaped facultative anaerobe with bipolar staining (giving it a safety pin appearance) causes the infection in mammals and humans.[1] The bacteria maintain their existence in a cycle involving rodents and their fleas. The genus Yersinia is gram-negative, bipolar staining coccobacilli, and, similarly to other Enterobacteriaceae, it has a fermentative metabolism. Y. pestis produces an antiphagocytic slime. The organism is motile when isolated, but becomes nonmotile in the mammalian host.
Taxonomy
Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacteriales; Yersinia; Yersinia pestis
Biology
 |
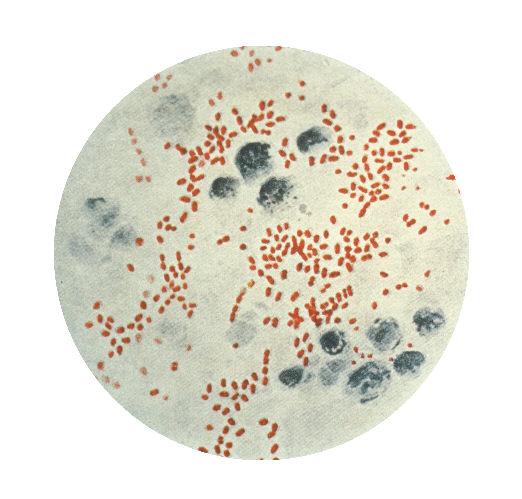 |
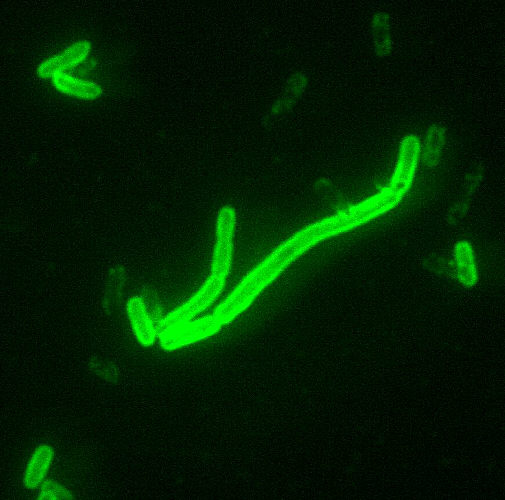
Yersinia pestis is a nonmotile, non-spore-forming, Gram-negative, non-lactose fermenting, ovoid and "safety-pin-shaped" (bipolar appearance when stained) bacillus. It is an enterobacteriaceae commonly measuring 0.75x1.5 μm. On cell cultures, it grows on sheep-blood agar, as grayish translucent colonies. It also grows on MacConkey and nutrient-rich broths.[4]
Yersinia pestis only survives for a few hours on physical surfaces, being very sensitive to high temperatures, sunlight and disinfectants.[4][5][6]
Yersinia pestis is thought to have evolved from Yersinia pseudotuberculosis about 1500 - 2000 years ago. According to its behavior towards nitrate and glycerol, Yersinia pestis may be classified as:[4]
- Biovar Antiqua
- Nitrate reduction: positive
- Glycerol reduction: positive
- Biovar Medievalis
- Nitrate reduction: negative
- Glycerol use: positive
- Biovar Orientalis
- Nitrate reduction: positive
- Glycerol use: negative
Genome
The complete genomic sequence is available for two of the three sub-species of yersinia pestis:
As of 2006, the genomic sequence of a strain of biovar Antiqua has been completed.[9] Similar to the other pathogenic strains, there are signs of mutations causing loss of function. The chromosome of strain KIM is 4,600,755 base pairs long; the chromosome of strain CO92 is 4,653,728 base pairs long.
Similarly to other enterobacteriaceae (Yersinia pseudotuberculosis and Yersinia enterocolitica), Yersinia pestis is host to the plasmid pCD1. In addition, it also hosts two other plasmids, pPCP1 (also called pPla or pPst) and pMT1 (also called pFra) that are not carried by the other Yersinia species.
- pFra codes for a phospholipase D that is important for the ability of Yersinia pestis to be transmitted by fleas.
- pPla codes for a protease, Pla, that activates plasminogen in human hosts and is a very important virulence factor for pneumonic plague.[10]
Together, these plasmids, and a pathogenicity island called HPI, encode several pathogenic proteins, characteristic of Yersinia pestis. Among other things, these virulence factors are required of:
- Bacterial adhesion
- Injection of proteins into the host cell
- Invasion of the host cell (via a Type III secretion system)
- Acquisition and binding of iron from red blood cell, via siderophores
A comprehensive and comparative proteomics analysis of Yersinia pestis strain KIM was performed in 2006.[11] The analysis focused on the transition to a growth condition mimicking growth in host cells.
Tropism
Yersinia pestis shows tropism for lymphoid tissue.
Natural Reservoir
Plague is primarily a disease of rodents. The infection is maintained in natural foci of the disease in wild rodent colonies, through transmission between rodents, by their flea ectoparasites.[12]
Gallery
-
Gram-negative Yersinia pestis bacteria, which was grown on a medium of sheep’s blood agar (SBA) 72hrs. From Public Health Image Library (PHIL). [13]
-
Ulcerated skin lesion at a plague inoculation site caused by the Gram-negative bacterium, Yersinia pestis. From Public Health Image Library (PHIL). [13]
-
Blood smear reveals presence of Gram-negative Yersinia pestis plague bacteria. From Public Health Image Library (PHIL). [13]
-
Gram-negative Yersinia pestis bacteria, which were cultured on a sheep blood agar (SBA) medium 120 hrs (20x mag). From Public Health Image Library (PHIL). [13]
-
Gram-negative Yersinia pestis bacteria, which were cultured on a sheep blood agar (SBA) medium 120 hrs (20x mag). From Public Health Image Library (PHIL). [13]
-
Gram-negative Yersinia pestis bacteria, which were cultured on a sheep blood agar (SBA) medium 24 hrs (10x mag). From Public Health Image Library (PHIL). [13]
-
Gram-negative Yersinia pestis bacteria, which were cultured on a sheep blood agar (SBA) medium 72 hrs (10x mag). From Public Health Image Library (PHIL). [13]
-
Hematoxylin-eosin stained (H&E) photomicrograph reveals histopathologic changes in a lymph node tissue sample in a case of fatal human plague. From Public Health Image Library (PHIL). [13]
-
Hematoxylin-eosin stained lung tissue sample revealed the histopathologic changes indicative of what was diagnosed as a case of fatal human plague from the country of Nepal (125x mag). From Public Health Image Library (PHIL). [13]
-
Chest x-ray of a plague patient revealing bilateral infection, greater on the patient's left side, which was diagnosed as a case of pneumonic plague, caused by Yersinia pestis. From Public Health Image Library (PHIL). [13]
-
Fingertips of patient’s right hand exhibited the signs of what is known as acral gangrene caused by the bacterium, Yersinia pestis. From Public Health Image Library (PHIL). [13]
-
Patient acquired a plague infection through abrasions on his upper right leg. From Public Health Image Library (PHIL). [13]
-
Lung tissue sample revealed the histopathologic changes indicative of fatal human plague (125x mag). From Public Health Image Library (PHIL). [13]
-
Purple-colored Yersinia pestis bacteria on the proventricular spines of a Xenopsylla cheopis flea. From Public Health Image Library (PHIL). [13]
References
- ↑ Collins FM (1996). Pasteurella, Yersinia, and Francisella. In: Baron's Medical Microbiology (Baron S et al, eds.) (4th ed.). Univ. of Texas Medical Branch. ISBN 0-9631172-1-1.
- ↑ "http://phil.cdc.gov/phil/details.asp". External link in
|title=(help) - ↑ "http://phil.cdc.gov/phil/details.asp". External link in
|title=(help) - ↑ 4.0 4.1 4.2 Koirala, Janak (2006). "Plague: Disease, Management, and Recognition of Act of Terrorism". Infectious Disease Clinics of North America. 20 (2): 273–287. doi:10.1016/j.idc.2006.02.004. ISSN 0891-5520.
- ↑ Perry RD, Fetherston JD (1997). "Yersinia pestis--etiologic agent of plague". Clin Microbiol Rev. 10 (1): 35–66. PMC 172914. PMID 8993858.
- ↑ Gage KL, Dennis DT, Orloski KA, Ettestad P, Brown TL, Reynolds PJ; et al. (2000). "Cases of cat-associated human plague in the Western US, 1977-1998". Clin Infect Dis. 30 (6): 893–900. doi:10.1086/313804. PMID 10852811.
- ↑ Deng W; et al. (2002). "Genome Sequence of Yersinia pestis KIM". Journal of Bacteriology. 184 (16): 4601&ndash, 4611. doi:10.1128/JB.184.16.4601-4611.2002. PMC 135232. PMID 12142430. Unknown parameter
|author-separator=ignored (help) - ↑ Parkhill J; et al. (2001). "Genome sequence of Yersinia pestis, the causative agent of plague". Nature. 413 (6855): 523&ndash, 527. doi:10.1038/35097083. PMID 11586360. Unknown parameter
|author-separator=ignored (help) - ↑ Chain PS; Hu P; Malfatti SA; et al. (2006). "Complete Genome Sequence of Yersinia pestis Strains Antiqua and Nepal516: Evidence of Gene Reduction in an Emerging Pathogen". J. Bacteriol. 188 (12): 4453–63. doi:10.1128/JB.00124-06. PMC 1482938. PMID 16740952. Unknown parameter
|author-separator=ignored (help) - ↑ Lathem WW, Price PA, Miller VL, Goldman WE (2007). "A plasminogen-activating protease specifically controls the development of primary pneumonic plague". Science. 315 (5811): 509–13. doi:10.1126/science.1137195. PMID 17255510.
- ↑ Hixson K; et al. (2006). "Biomarker candidate identification in Yersinia pestis using organism-wide semiquantitative proteomics". Journal of Proteome Research. 5 (11): 3008–3017. doi:10.1021/pr060179y. PMID 16684765. Unknown parameter
|author-separator=ignored (help) - ↑ "Plague".
- ↑ 13.00 13.01 13.02 13.03 13.04 13.05 13.06 13.07 13.08 13.09 13.10 13.11 13.12 13.13 "Public Health Image Library (PHIL)".
![Gram-negative Yersinia pestis bacteria, which was grown on a medium of sheep’s blood agar (SBA) 72hrs. From Public Health Image Library (PHIL). [13]](/images/6/62/Enterobacteria34.jpeg)
![Ulcerated skin lesion at a plague inoculation site caused by the Gram-negative bacterium, Yersinia pestis. From Public Health Image Library (PHIL). [13]](/images/e/ef/Enterobacteria31.png)
![Blood smear reveals presence of Gram-negative Yersinia pestis plague bacteria. From Public Health Image Library (PHIL). [13]](/images/2/26/Enterobacteria29.jpeg)
![Gram-negative Yersinia pestis bacteria, which were cultured on a sheep blood agar (SBA) medium 120 hrs (20x mag). From Public Health Image Library (PHIL). [13]](/images/c/cf/Enterobacteria22.jpeg)
![Gram-negative Yersinia pestis bacteria, which were cultured on a sheep blood agar (SBA) medium 120 hrs (20x mag). From Public Health Image Library (PHIL). [13]](/images/a/a2/Enterobacteria21.jpeg)
![Gram-negative Yersinia pestis bacteria, which were cultured on a sheep blood agar (SBA) medium 24 hrs (10x mag). From Public Health Image Library (PHIL). [13]](/images/c/c3/Enterobacteria20.jpeg)
![Gram-negative Yersinia pestis bacteria, which were cultured on a sheep blood agar (SBA) medium 72 hrs (10x mag). From Public Health Image Library (PHIL). [13]](/images/c/c5/Enterobacteria19.jpeg)
![Hematoxylin-eosin stained (H&E) photomicrograph reveals histopathologic changes in a lymph node tissue sample in a case of fatal human plague. From Public Health Image Library (PHIL). [13]](/images/9/9b/Bubonic_plague16.jpeg)
![Hematoxylin-eosin stained lung tissue sample revealed the histopathologic changes indicative of what was diagnosed as a case of fatal human plague from the country of Nepal (125x mag). From Public Health Image Library (PHIL). [13]](/images/1/11/Bubonic_plague05.jpeg)
![Chest x-ray of a plague patient revealing bilateral infection, greater on the patient's left side, which was diagnosed as a case of pneumonic plague, caused by Yersinia pestis. From Public Health Image Library (PHIL). [13]](/images/6/63/Bubonic_plague15.jpeg)
![Fingertips of patient’s right hand exhibited the signs of what is known as acral gangrene caused by the bacterium, Yersinia pestis. From Public Health Image Library (PHIL). [13]](/images/2/27/Bubonic_plague14.jpeg)
![Patient acquired a plague infection through abrasions on his upper right leg. From Public Health Image Library (PHIL). [13]](/images/a/af/Bubonic_plague12.jpeg)
![Lung tissue sample revealed the histopathologic changes indicative of fatal human plague (125x mag). From Public Health Image Library (PHIL). [13]](/images/1/1c/Bubonic_plague04.jpeg)
![Purple-colored Yersinia pestis bacteria on the proventricular spines of a Xenopsylla cheopis flea. From Public Health Image Library (PHIL). [13]](/images/2/2b/Bubonic_plague01.jpeg)