Polyploidy
|
WikiDoc Resources for Polyploidy |
|
Articles |
|---|
|
Most recent articles on Polyploidy |
|
Media |
|
Evidence Based Medicine |
|
Clinical Trials |
|
Ongoing Trials on Polyploidy at Clinical Trials.gov Clinical Trials on Polyploidy at Google
|
|
Guidelines / Policies / Govt |
|
US National Guidelines Clearinghouse on Polyploidy
|
|
Books |
|
News |
|
Commentary |
|
Definitions |
|
Patient Resources / Community |
|
Patient resources on Polyploidy Discussion groups on Polyploidy Patient Handouts on Polyploidy Directions to Hospitals Treating Polyploidy Risk calculators and risk factors for Polyploidy
|
|
Healthcare Provider Resources |
|
Causes & Risk Factors for Polyploidy |
|
Continuing Medical Education (CME) |
|
International |
|
|
|
Business |
|
Experimental / Informatics |
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [1]
Please Take Over This Page and Apply to be Editor-In-Chief for this topic: There can be one or more than one Editor-In-Chief. You may also apply to be an Associate Editor-In-Chief of one of the subtopics below. Please mail us [2] to indicate your interest in serving either as an Editor-In-Chief of the entire topic or as an Associate Editor-In-Chief for a subtopic. Please be sure to attach your CV and or biographical sketch.
Overview
Polyploidy occurs in cells and organisms when there are more than two homologous sets of chromosomes. Polyploid types are labelled according to the number of chromosome sets in the nucleus:
- triploid (three sets; 3x), for example the genus Tardigrada
- tetraploid (four sets; 4x), for example Salmonidae fish
- pentaploid (five sets; 5x)
- hexaploid (six sets; 6x), for example wheat
- oktoploid (eight sets; 8x), for example Acipenser (genus of sturgeon fish)
- dekaploid (ten sets; 10x), for example certain strawberries
- dodecaploid (twelve sets; 12x), for example the plant Celosia argentea
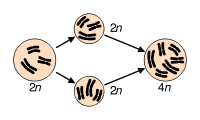
Most organisms are normally diploid; polyploidy may occur due to abnormal cell division. It is most commonly found in plants. Haploidy may also occur as a normal stage in an organism's life. A haploid has only one set of chromosomes.
Polyploidy occurs in some animals, such as goldfish, salmon, and salamanders, but is especially common among ferns and flowering plants, including both wild and cultivated species. Wheat, for example, after millennia of hybridization and modification by humans, has strains that are diploid (two sets of chromosomes), tetraploid (four sets of chromosomes) with the common name of durum or macaroni wheat, and hexaploid (six sets of chromosomes) with the common name of bread wheat. Many agriculturally important plants of the genus Brassica are also tetraploids; their relationship is described by the Triangle of U.
The occurrence of polyploidy is a mechanism of speciation and is known to have resulted in new species of the plant Salsify (also known as "goatsbeard").
Examples in animals are more common in the ‘lower’ forms such as flatworms, leeches, and brine shrimp. Polyploid animals are often sterile, so they often reproduce by parthenogenesis. Polyploid salamanders and lizards are also quite common and parthenogenetic. While mammalian liver cells are polyploid, rare instances of polyploid mammals are known, but most often result in prenatal death.
The only known exception to this rule is an octodontid rodent of Argentina's harsh desert regions, known as the Red Viscacha-Rat (Tympanoctomys barrerae). This rodent is not a rat, but kin to [uinea pigs and chinchillas. Its "new" diploid [2n] number is 102 and so its cells are roughly twice normal size. Its closest living relation is Octomys mimax, the Andean Viscacha-Rat of the same family, whose 2n=56. It is surmised that an Octomys-like ancestor produced tetraploid (i.e., 4n=112) offspring that were, by virtue of their doubled chromosomes, reproductively isolated from their parents; but that these likely survived the ordinarily catastrophic effects of polyploidy in mammals by shedding (via translocation or some similar mechanism) the "extra" set of sex chromosomes gained at this doubling.
Polyploidy can be induced in cell culture by some chemicals: the best known is colchicine, which can result in chromosome doubling, though its use may have other less obvious consequences as well.
There are large number of polyploid crop varieties - See Polyploid Crops below.
There are few naturally occurring polyploid conifers. One example is the giant tree Sequoia sempervirens or Coast Redwood which is a hexaploid (6x) with 66 chromosomes (2n=6x=66), although the origin is unclear [1].
Polyploidy in humans (Aneuploidy)
True polyploidy rarely occurs in humans, although it occurs in some tissues (especially in the liver). Polyploidy refers to a numerical change in a whole set of chromosomes. Organisms in which a particular chromosome, or chromosome segment, is under- or overrepresented are said to be aneuploid (from the Greek words meaning "not," "good," and "fold"). Therefore the distinction between aneuploidy and polyploidy is that aneuploidy refers to a numerical change in part of the chromosome, whereas polyploidy refers to a numerical change in the whole set of chromosomes. [2]: Cytogenetic Variation (p109)]
Aneuploidy occurs in humans in the form of triploidy (69,XXX) and tetraploidy (92,XXXX), not to be confused with 47,XXX or 48, XXXX aneuploidy. Triploidy, usually due to polyspermy, occurs in about 2-3% of all human pregnancies and ~15% of miscarriages. The vast majority of triploid conceptions end as miscarriage and those that do survive to term typically die shortly after birth. In some cases survival past birth may occur longer if there is mixoploidy with both a diploid and a triploid cell population present.
Triploidy may be the result of either digyny (the extra haploid set is from the mother) or diandry (the extra haploid set is from the father). Diandry is almost always caused by the fertilization of an egg by two sperm (dispermy). Digyny is most commonly caused by either failure of one meiotic division during oogenesis leading to a diploid oocyte or failure to extrude one polar body from the oocyte. Diandry appears to predominate among early miscarriages while digyny predominates among triploidy that survives into the fetal period. However, among early miscarriages, digyny is also more common in those cases <8.5 weeks gestational age or those in which an embryo is present. There are also two distinct phenotypes in triploid placentas and fetuses that are dependent on the origin of the extra haploid set. In digyny there is typically an asymmetric poorly grown fetus, with marked adrenal hypoplasia and a very small placenta. In diandry, the fetus (when present) is typically normally grown or symmetrically growth restricted, with normal adrenal glands and an abnormally large cystic placenta that is called a partial hydatidiform mole. These parent-of-origin effects reflect the effects of genomic imprinting.
Complete tetraploidy is more rarely diagnosed than triploidy, but is observed in 1-2% of early miscarriages. However, some tetraploid cells are not uncommonly found in chromosome analysis at prenatal diagnosis and these are generally considered ‘harmless’. It is not clear whether these tetraploid cells simply tend to arise during in vitro cell culture or whether they are also present in placental cells in vivo. There are, at any rate, very few clinical reports of fetuses/infants diagnosed with tetraploidy mosaicism.
Mixoploidy is quite commonly observed in human preimplantation embryos and includes haploid/diploid as well as diploid/tetraploid mixed cell populations. It is unknown whether these embryos fail to implant and are therefore rarely detected in ongoing pregnancies or if there is simply a selective process favoring the diploid cells.
Polyploidy in plants
Polyploidy is pervasive in plants and some estimates suggest that 30-80% of living plant species are polyploid, and many lineages show evidence of ancient polyploidy (paleopolyploidy) in their genomes.[3] Huge explosions in angiosperm species diversity appear to have coincided with the timing of ancient genome duplications shared by many species.[4] Polyploid plants can arise spontaneously in nature by several mechanisms, including meiotic or mitotic failures, and fusion of unreduced (2n) gametes.[5] Both autopolyploids (eg. potato) and allopolyploids (eg. canola, wheat, cotton) can be found among both wild and domesticated plant species. Most polyploids display heterosis relative to their parental species, and may display novel variation or morphologies that may contribute to the processes of speciation and eco-niche exploitation.[6] The mechanisms leading to novel variation in newly formed allopolyploids may include gene dosage effects (resulting from more numerous copies of genome content), the reunion of divergent gene regulatory hierarchies, chromosomal rearrangements, and epigenetic remodeling, all of which affect gene content and/or expression levels.[7] Many of these rapid changes may contribute to reproductive isolation and speciation.
Polyploid crops
Polyploid plants tend to be larger and better at flourishing in early succession habitats such as farm fields. In the breeding of crops, the tallest and best thriving plants are selected for. Thus, many crops (and agricultural weeds) may have unintentionally been bred to a higher level of ploidy.
The induction of polyploids is a common technique to overcome the sterility of a hybrid species during plant breeding. For example, Triticale is the hybrid of wheat (Triticum turgidum) and rye (Secale cereale). It combines sought-after characteristics of the parents, but the initial hybrids are sterile. After polyploidization, the hybrid becomes fertile and can thus be further propagated to become triticale.
In some situations polyploid crops are preferred because they are sterile. For example many seedless fruit varieties are seedless as a result of polyploidy. Such crops are propagated using asexual techniques such as grafting.
Polyploidy in crop plants is most commonly induced by treating seeds with the chemical colchicine.
Examples of Polyploid Crops
- Triploid crops: banana, apple, ginger, watermelon, citrus[8]
- Tetraploid crops: durum or macaroni wheat, maize, cotton, potato, cabbage, leek, tobacco, peanut, kinnow, Pelargonium
- Hexaploid crops: chrysanthemum, bread wheat, triticale, oat
- Octaploid crops: strawberry, dahlia, pansies, sugar cane
Some crops are found in a variety of ploidy. Apples, tulips and lilies are commonly found as both diploid and as triploid. Daylilies (Hemerocallis) cultivars are available as either diploid or tetraploid. Kinnows can be tetraploid, diploid, or triploid.
Terminology
Autopolyploidy
Autopolyploids are polyploids with multiple chromosome sets derived from a single species. Autopolyploids can arise from a spontaneous, naturally occurring genome doubling (for example, the potato). Others might form following fusion of 2n gametes unreduced gametes). Bananas and apples can be found as triploid autopolyploids. Autopolyploid plants typically display polysomic inheritance, and are therefore often infertile and propagated clonally
Allopolyploidy
Allopolyploids are polyploids with chromosomes derived from different species. Triticale is an example of an allopolyploid, having six chromosome sets, four from wheat (Triticum turgidum) and two from rye (Secale cereale). Amphidiploid is another word for an allopolyploid. Some of the best examples of allopolyploids come from the Brassicas, and the Triangle of U describes the relationships among the three common diploid Brassicas (B. oleracea, B. rapa, and B. nigra) and three allotetraploids (B. napus, B. juncea, and B. carinata) derived from hybridization among the diploids.
Homoeologous
The term is used to describe the relationship of similar chromosomes or parts of chromosomes brought together following inter-species hybridization and allopolyploidization, and whose relationship was completely homologous in an ancestral species. In allopolyploids, the homologous chromosomes within each parental sub-genome should pair faithfully during meiosis, leading to disomic inheritance; however in some allopolyploids, the homoeologous chromosomes of the parental genomes may be nearly as similar to one another as the homolgous chromosomes, leading to tetrasomic inheritance (four chromosomes pairing at meiosis), intergenomic recombination, and reduced fertility.
Homologous
The term is used to describe the relationship of similar chromosomes that pair at mitosis and meiosis. In a diploid, one homolog is derived from the male parent (pollen or sperm) and one is derived from the female parent (egg). During meiosis and gametogenesis, homologous chromosomes pair and exchange genetic material by recombination, leading to the production of sperm/pollen or eggs with chromosome haplotypes containing novel genetic variation.
Karyotype
A karyotype is the characteristic chromosome complement of a eukaryote species.[9][10] The preparation and study of karyotypes is part of cytology and, more specifically, cytogenetics.
Although the replication and transcription of DNA is highly standardized in eukaryotes, the same cannot be said for their karotypes, which are highly variable between species in chromosome number and in detailed organization despite being constructed out of the same macromolecules. In some cases there is even significant variation within species. This variation provides the basis for a range of studies in what might be called evolutionary cytology.
Paralogous
The term is used to describe the relationship among duplicated genes or portions of chromosomes that derived from a common ancestral DNA. Paralogous segments of DNA may arise spontaneously by errors during DNA replication, copy and paste transposons, or whole genome duplications.
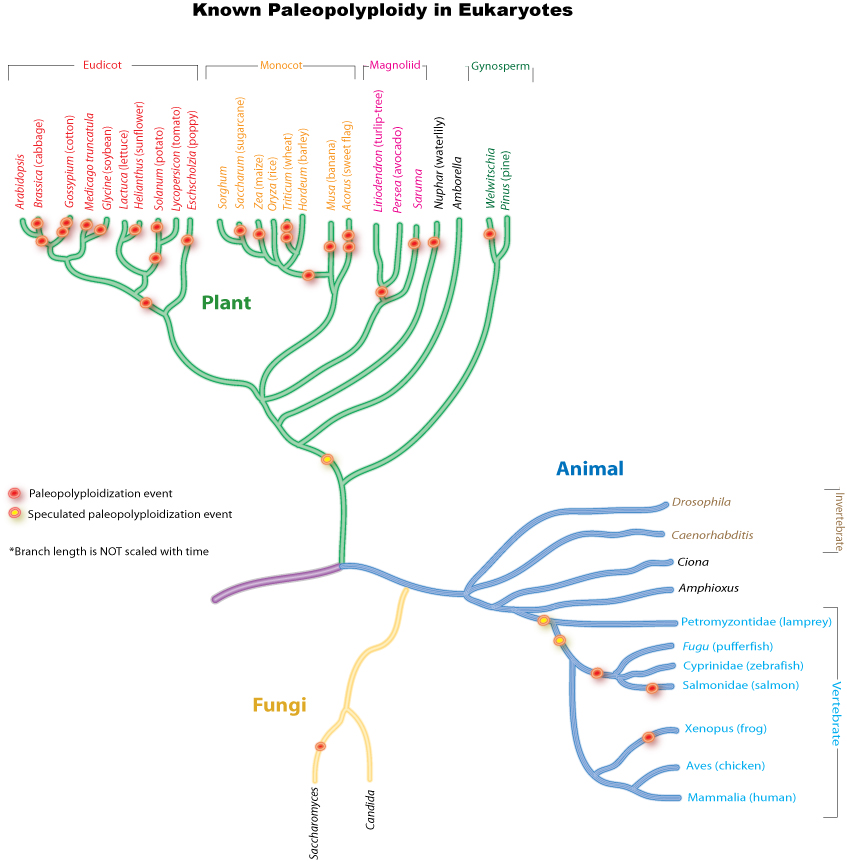
Paleopolyploidy
Ancient genome duplications probably occurred in the evolutionary history of all life. Duplication events that occurred long ago in the history of various evolutionary lineages can be difficult to detect because of subsequent diploidization (such that a polyploid starts to behave cytogenetically as a diploid over time) as mutations and gene translations gradually make one copy of each chromosome unlike its other copy.
In many cases, these events can be inferred only through comparing sequenced genomes. Examples of unexpected but recently confirmed ancient genome duplications include the baker's yeast (Saccharomyces cerevisiae), mustard weed/thale cress (Arabidopsis thaliana), rice (Oryza sativa), and an early evolutionary ancestor of the vertebrates (which includes the human lineage) and another near the origin of the teleost fishes. Angiosperms (flowering plants) have paleopolyploidy in their ancestry. All eukaryotes probably have experienced a polyploidy event at some point in their evolutionary history.
See also
References
- ↑ Ahuja MR, Neale DB. "Origins of Polyploidy in Coast Redwood (Sequoia sempervirens (D. DON) ENDL.) and Relationship of Coast Redwood to other Genera of Taxodiaceae" Silvae Genetica 51, 2–3 (2002)
- ↑ Griffiths, A. J. et al. 2000. An introduction to genetic analysis, 7th ed. W. H. Freeman, New York ISBN 0-7167-3520-2
- ↑ Meyers and Levin 2006; Rieseberg and Willis 2007; Otto 2007
- ↑ de Bodt et al 2005
- ↑ Comai 2005
- ↑ Comai 2005; Rieseberg and Willis 2007
- ↑ Osborn et al., 2003; Chen and Ni 2006; Chen 2007
- ↑ Seedless Fruits Make Others Needless
- ↑ White M.J.D. 1973. The chromosomes. 6th ed, Chapman & Hall, London. p28
- ↑ Stebbins G.L. 1950. Variation and evolution in plants. Chapter XII: The Karyotype. Columbia University Press N.Y.
Further reading
- Snustad, P. et al. 2006. Principles of Genetics, 4th ed. John Wiley & Sons, Inc. Hoboken, NJ ISBN 10 0-471-69939-X
- Arabidopsis Genome Initiative (2000). Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408: 796-815.
- Eakin, G.S. & Behringer, R.R. (2003). Tetraploid development in the mouse. Developmental Dynamics 228: 751-766.
- Gaeta, R.T., Pires, J.C., Iniguez, F.L., Leon, E., and Osborn, T.C. (2007). Genomic changes in resynthesized Brassica napus and their effect on gene expression and phenotype. "Plant Cell" PMID: 18024568.
- Gregory, T.R. & Mable, B.K. (2005). Polyploidy in animals. In The Evolution of the Genome (edited by T.R. Gregory). Elsevier, San Diego, pp. 427-517.
- Jaillon, O. et al. (2004). Genome duplication in the teleost fish Tetraodon nigroviridis reveals the early vertebrate proto-karyotype. Nature 431: 946-957.
- Paterson, A.H., Bowers, J. E., Van de Peer, Y. & Vandepoele, K. (2005). Ancient duplication of cereal genomes. New Phytologist 165: 658-661.
- Raes, J., Vandepoele, K., Saeys, Y., Simillion, C. & Van de Peer, Y. (2003). Investigating ancient duplication events in the Arabidopsis genome. Journal of Structural and Functional Genomics 3: 117-129.
- Simillion, C., Vandepoele, K., Van Montagu, M., Zabeau, M. & Van de Peer, Y. (2002). The hidden duplication past of Arabidopsis thaliana. Proceedings of the National Academy of Science of the USA 99: 13627-13632.
- Soltis, D. E.; Soltis, P. S.; Schemske, D. W.; Hancock, J. F.; Thompson, J. N.; Husband, B. C. & Judd, W. S. (2007).Autopolyploidy in angiosperms: have we grossly underestimated the number of species? Taxon 56 (1):13-30.
- Taylor, J.S., Braasch, I., Frickey, T., Meyer, A. & Van de Peer, Y. (2003). Genome duplication, a trait shared by 22,000 species of ray-finned fish. Genome Research 13: 382-390.
- Tate, J.A., Soltis, D.E., & Soltis, P.S. (2005). Polyploidy in plants. In The Evolution of the Genome (edited by T.R. Gregory). Elsevier, San Diego, pp.371-426.
- Van de Peer, Y., Taylor, J.S. & Meyer, A. (2003). Are all fishes ancient polyploids? Journal of Structural and Functional Genomics 3: 65-73.
- Van de Peer, Y. (2004). Tetraodon genome confirms Takifugu findings: most fish are ancient polyploids. Genome Biology 5(12):250.
- Van de Peer, Y. and Meyer, A. (2005). Large-scale gene and ancient genome duplications. In The Evolution of the Genome (edited by T.R. Gregory). Elsevier, San Diego, pp.329-368
- Wolfe, K.H. & Shields, D.C. (1997). Molecular evidence for an ancient duplication of the entire yeast genome. Nature 387: 708-713.
- Wolfe, K.H. (2001). Yesterday's polyploids and the mystery of diploidization. Nature Reviews Genetics 2: 333-341.
External links
- Polyploidy on Kimball's Biology Pages
- The polyploidy portal a community-editable project with information, research, education, and a bibliography about polyploidy.
Template:SIB Template:Speciation
da:Polyploidi de:Polyploidie id:Poliploidi lv:Poliploīdija nl:Polyploïdie fi:Polyploidia sv:Polyploidi