Chemotaxis
Overview
Chemotaxis, a kind of taxis, is the phenomenon in which bodily cells, bacteria, and other single-cell or multicellular organisms direct their movements according to certain chemicals in their environment. This is important for bacteria to find food (for example, glucose) by swimming towards the highest concentration of food molecules, or to flee from poisons (for example, phenol). In multicellular organisms, chemotaxis is critical to development as well as normal function. In addition, it has been recognized that mechanisms that allow chemotaxis in animals can be subverted during cancer metastasis.
Chemotaxis is called positive if movement is in the direction of a higher concentration of the chemical in question, and negative if the direction is opposite.
History of chemotaxis research

Although migration of cells was detected from the early days of the development of microscopy (Leeuwenhoek), erudite description of chemotaxis was first made by T.W. Engelmann (1881) and W.F. Pfeffer (1884) in bacteria and H.S. Jennings (1906) in ciliates. The Nobel Prize Laureate E. Metchnikoff also contributed to the study of the field with investigations of the process as an initial step of phagocytosis. The significance of chemotaxis in biology and clinical pathology was widely accepted in the 1930s. The most fundamental definitions belonging to the phenomenon were also drafted by this time. The most important aspects in quality control of chemotaxis assays were described by H. Harris in the 1950s. In the 1960s and 1970s, the revolution of modern cell biology and biochemistry provided a series of novel techniques which became available to investigate the migratory responder cells and subcellular fractions responsible for chemotactic activity. The pioneering works of J. Adler represented a significant turning point in understanding the whole process of intracellular signal transduction of bacteria.[1]
On November 3, 2006, Dr. Dennis Bray of University of Cambridge was awarded the Microsoft European Science Award for his work on chemotaxis on E. coli.[2][3]
Phylogeny and chemotactic signalling
Chemotaxis is one of the most basic cell physiological responses. Development of receptor systems for the detection of harmful and favorable substances in the environment was most essential to unicellular organisms from the very early stages of phylogeny. Comprehensive analysis of chemotactic activity of the eukaryotic protozoon Tetrahymena pyriformis and consensus sequences of appearance of amino acids in the primordial soup suggest that there was a good correlation between the chemotactic character of these relative simple organic molecules and their development on the Earth. In this way the earliest molecules are suggested to be highly chemoattractant (e.g. Gly, Glu, Pro), while latter ones are thought to be strongly chemorepellent (e.g. Tyr, Trp, Phe) amino acids.[4]
Bacterial chemotaxis
Some bacteria, such as E. coli, have several flagella per cell (4–10 typically). These can rotate in two ways :
- Counter-clockwise rotation aligns the flagella into a single rotating bundle, causing the bacterium to swim in a straight line.
- Clockwise rotation breaks the flagella bundle apart such that each flagellum points in a different direction, causing the bacterium to tumble in place.
The directions of rotation are given for an observer outside the cell looking down the flagella toward the cell.

Behavior
The overall movement of a bacterium is the result of alternating tumble and swim phases. If one watches a bacterium swimming in a uniform environment, its movement will look like a random walk with relatively straight swims interrupted by random tumbles that reorient the bacterium. Bacteria such as E. coli are unable to choose the direction in which they swim, and are unable to swim in a straight line for more than a few seconds due to rotational diffusion. In other words, bacteria "forget" the direction in which they are going. Given these limitations, it is remarkable that bacteria can direct their motion to find favorable locations with high concentrations of attractants (usually food) and avoid repellents (usually poisons).
In the presence of a chemical gradient bacteria will chemotax, or direct their overall motion based on the gradient. If the bacterium senses that it is moving in the correct direction (toward attractant/away from repellent), it will keep swimming in a straight line for a longer time before tumbling. If it is moving in the wrong direction, it will tumble sooner and try a new direction at random. In other words, bacteria like E. coli use temporal sensing to decide whether life is getting better or worse. In this way, it finds the location with the highest concentration of attractant (usually the source) quite well. Even under very high concentrations, it can still distinguish very small differences in concentration. Fleeing from a repellent works with the same efficiency.
It seems remarkable that this purposeful random walk is a result of simply choosing between two methods of random movement; namely tumbling and straight swimming. In fact, chemotactic responses such as forgetting direction and choosing movements resemble the decision-making abilities of higher lifeforms with brains that process sensory data.
The helical nature of the individual flagellar filament is critical for this movement to occur. As such, the protein that makes up the flagellar filament, flagellin, is quite similar among all flagellated bacteria. Vertebrates seem to have taken advantage of this fact by possessing an immune receptor (TLR5) designed to recognize this conserved protein.
As in many instances in biology, there are bacteria that do not follow this rule. Many bacteria, such as Vibrio, are monoflagellated and have a single flagellum at one pole of the cell. Their method of chemotaxis is different. Others possess a single flagellum that is kept inside the cell wall. These bacteria move by spinning the whole cell, which is shaped like a corkscrew.[5]
Signal transduction
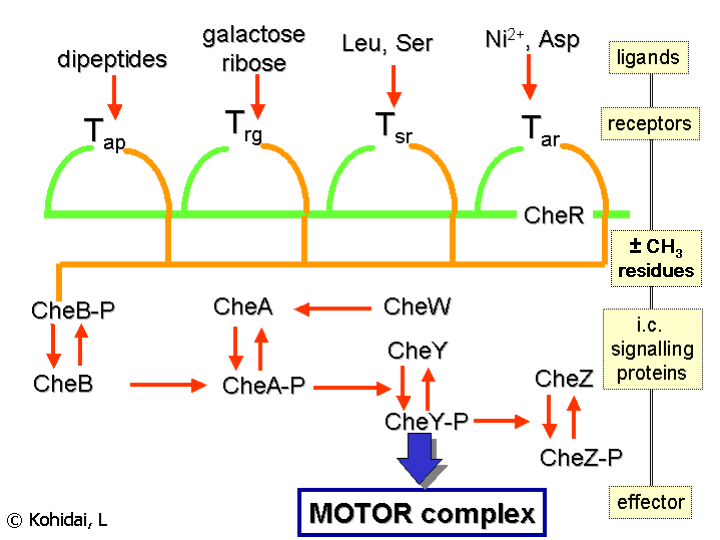
Chemical gradients are sensed through multiple transmembrane receptors, called methyl accepting chemotaxis proteins (MCPs), which vary in the molecules that they detect. These receptors may bind attractants or repellents directly or indirectly through interaction with proteins of periplasmatic space. The signals from these receptors are transmitted across the plasma membrane into the cytosol, where Che proteins are activated. The Che proteins alter the tumbling frequency, and alter the receptors.

Flagellum regulation
The proteins CheW and CheA bind to the receptor. The activation of the receptor by an external stimulus causes autophosphorylation in the histidine kinase, CheA, at a single highly conserved histidine residue. CheA in turn transfers phosphoryl groups to conserved aspartate residues in the response regulators CheB and CheY [ note: CheA is a histidine kinase and it does not actively transfer the phosphoryl group. The response regulator CheB takes the phosphoryl group from CheA]. This mechanism of signal transduction is called a 'Two Component System' and is a common form of signal transduction in bacteria. CheY induces tumbling by interacting with the flagellar switch protein FliM, inducing a change from counter-clockwise to clockwise rotation of the flagellum. Change in the rotation state of a single flagellum can disrupt the entire flagella bundle and cause a tumble.
Receptor regulation
CheB, when activated by CheA, acts as a methylesterase, removing methyl groups from glutamate residues on the cytosolic side of the receptor. It works antagonistically with CheR, a methyltransferase, which adds methyl residues to the same glutamate residues. The more methyl residues are attached to the receptor, the more sensitive the receptor. As the signal from the receptor induces demethylation of the receptor in a feedback loop, the system is continuously adjusted to environmental chemical levels, remaining sensitive for small changes even under extreme chemical concentrations. This regulation allows the bacterium to 'remember' chemical concentrations from the recent past and compare them to those it is currently experiencing, thus 'know' whether it is traveling up or down a gradient. However, the methylation system alone cannot account for the wide range of sensitivity that bacteria have to chemical gradients. Additional regulatory mechanisms such as receptor clustering and receptor-receptor interactions also modulate the signalling pathway.
Eukaryotic chemotaxis
The mechanism by which eukaryotic cells chemotax is quite different from that in bacteria; however, sensing of chemical gradients is still a crucial step in the process. Due to their size, prokaryotes cannot detect effective concentration gradients, therefore these cells scan and evaluate their environment by a constant swimming (consecutive steps of straight swims and tumbles). In contrast to prokaryotes, the size of eukaryotic cells allows for the possibility of detecting gradients, which results in a dynamic and polarized distribution of receptors. Induction of these receptors by chemoattractants or chemorepellents results in migration towards or away from the chemotactic substance.

Levels of receptors, intracellular signalling pathways and the effector mechanisms all represent diverse, eukaryotic type components. In eukaryotic unicellular cells, ameboid movement and cilium or the eukaryotic flagellum are the main effectors (e.g. Amoeba or Tetrahymena). Some eukaryotic cells of higher vertebrate origin, such as immune cells also move to where they need to be. Besides immune competent cells (granulocyte, monocyte, lymphocyte) a large group of cells - considered previously to be fixed into tissues - are also motile in special physiological (e.g. mast cell, fibroblast, endothelial cells)or pathological conditions (e.g. metastases). Chemotaxis has high significance in the early phases of embryogenesis as development of germ layers is guided by gradients of signal molecules.
Motility
Unlike motility in bacterial chemotaxis, the mechanism by which eukaryotic cells physically move is unclear. There appear to be mechanisms by which an external chemotactic gradient is sensed and turned into an intracellular PIP3 gradient, which results in a gradient in the activation of signaling pathway culminating in the polymerisation of actin filaments. The growing distal end of actin filaments develops connections with the internal surface of the plasma membrane via different sets of peptides and results in the formation of pseudopods. Cilium of eukaryotic cells can also result in chemotaxis, while in this case it is mainly a Ca2+ dependent induction of the microtubular system of the basal body and the 9x2+2 microtubules stroke of cilia. The orchestered beating of hundreds of cilia is synchronized by a submembranous system built between basal bodies. The details of the signaling pathways are still not totally clear.
Chemotaxis related migratory responses
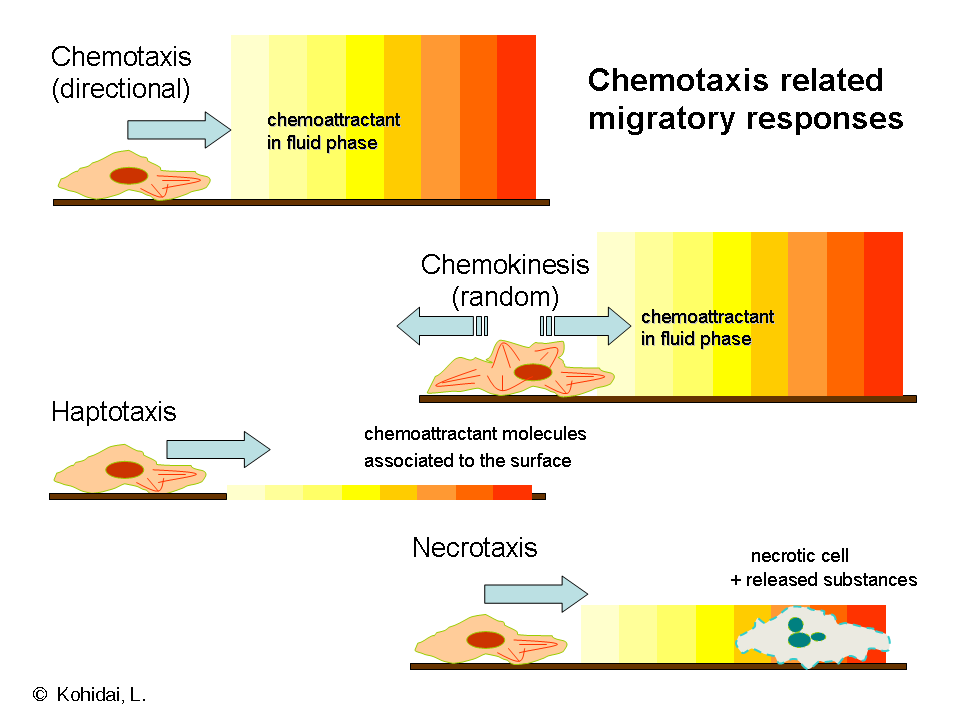
Although chemotaxis is the most frequently studied form of migration there are several other forms of locomotion on the cellular level.
- Chemokinesis is also induced by molecules of the liquid phase of the surrounding environment; however, the response elicited is a not vectorial, random taxis. Neither amplitude nor frequency of motion has characteristic, directional components as this behaviour provides more scanning of the environment than migration between two distinct points.
- In haptotaxis the gradient of the chemoattractant is expressed or bound on a surface, in contrast to the classical way of chemotaxis when the gradient develops in a soluble space. The main biologically active haptotactic surface is the extracellular matrix (ECM); the presence of bound ligands is responsible for induction of transendothelial migration and angiogenesis.
- Necrotaxis embodies a special type of chemotaxis when the chemoattractant molecules are released from necrotic or apoptotic cells. Depending on the chemical character of released substances necrotaxis can accumulate or repel cells, which underlines the pathophysiological significance of this phenomenon.

Receptors
For the most part, eukaryotic cells sense the presence of chemotactic stimuli though the use of 7-transmembrane (or serpentine) heterotrimeric G-protein coupled receptors. This class of receptors is huge, representing a significant portion of the genome. Some members of this gene superfamily are used in eyesight (rhodopsins) as well as in olfaction (smelling). The main classes of professional chemotaxis receptors are triggered by formyl peptides - formyl peptide receptors (FPR), chemokines - chemokine receptors (CCR or CXCR) and leukotrienes - leukotriene receptors (BLT); however, induction of a wide set of membrane receptors (e.g. amino acids, insulin, vasoactive peptides) also elicit migration of the cell.
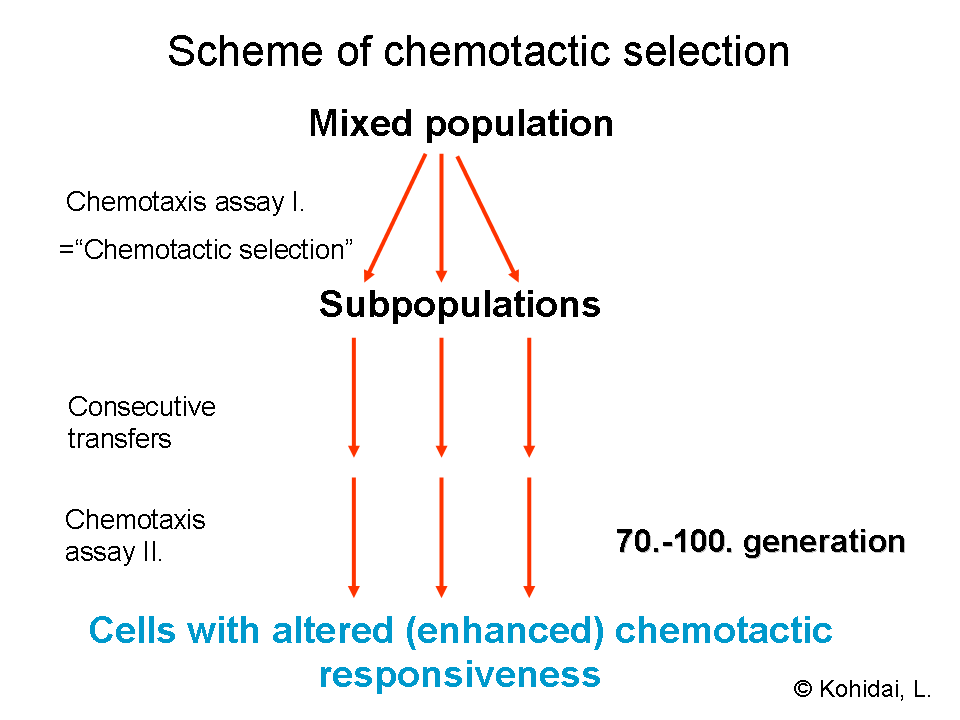
Chemotactic selection
While some chemotaxis receptors are expressed in the surface membrane with long-term characteristics as they are determined genetically, others have short-term dynamics as they are assembled ad hoc in the presence of the ligand. The diverse features of the chemotaxis receptors and ligands allows for the possibility of selecting chemotactic responder cells with a simple chemotaxis assay. By chemotactic selection we can determine whether a still uncharacterized molecule acts via the long- or the short-term receptor pathway. The term chemotactic selection is also used to designate a technique which separates eukaryotic or prokaryotic cells according to their chemotactic responsiveness to selector ligands.[6]
Chemotactic ligands
The number of molecules capable of eliciting chemotactic responses is relatively high, and we can distinguish primary and secondary chemotactic molecules. The main groups of the primary ligands are as follows:
- Formyl peptides are di-, tri-, tetrapeptides of bacterial origin (see formyl group on the N terminus of the peptide). They are released from bacteria in vivo or after decomposition of the cell. A typical member of this group is the N-formylmethionyl-leucyl-phenylalanine (fMLF or fMLP in references). The bacterial origin fMLF as a key component of inflammation has characteristic chemoattractant effects in neutrophil granulocytes and monocytes.
- Complement 3a (C3a) and complement 5a (C5a) are intermediate products of complement cascade. Their synthesis is joined to the three alternative pathways (classical, lectin dependent and alternative) of complement activation by a convertase enzyme. The main target cells of these derivaties are neutrophil garnulocytes and monocytes as well.
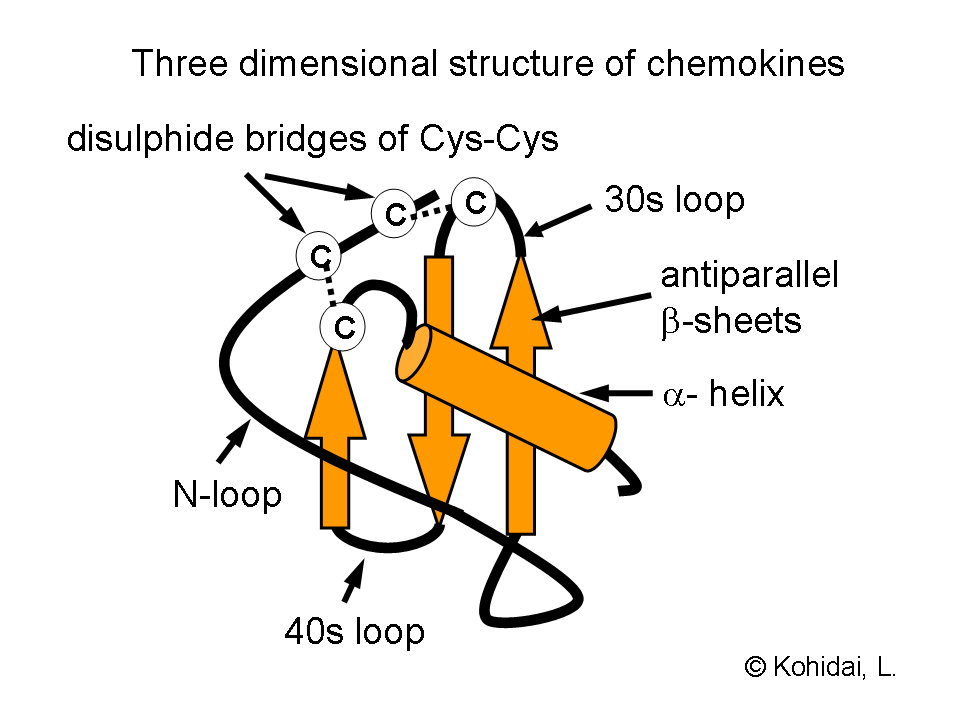
- Chemokines belong to a special class of cytokines. Their groups (C, CC, CXC, CX3C chemokines) represent not only structurally related molecules with a special arrangement of disulfide bridges, but their target cell specificity is also diverse: CC chemokines are acting on monocytes (e.g. RANTES), CXC chemokines are neutrophil granulocyte specific (e.g. IL-8).
Investigations of the three-dimensional structures of chemokines proved that a characteristic composition of beta-sheets and an alpha helix provides expression of sequences required for interaction with the chemokine receptors. Formation of dimers and their increased biological acitvity was demonstrated by crystallography of several chemokines e.g. IL-8.
- Leukotrienes belong to the group eicosanoids. They are significant lipid mediators of the arachidonic acid cascade converted by 5-lipoxigenase. Their predominant member is leukotriene B4 (LTB4) which elicits adhesion, chemotaxis and aggregation of leukocytes. The characteristic chemoattractant effect of LTB4 is induced via G-protein linked seven-transmembrane spanning leukotriene receptors which are highly expressed in inflammation and allergy.
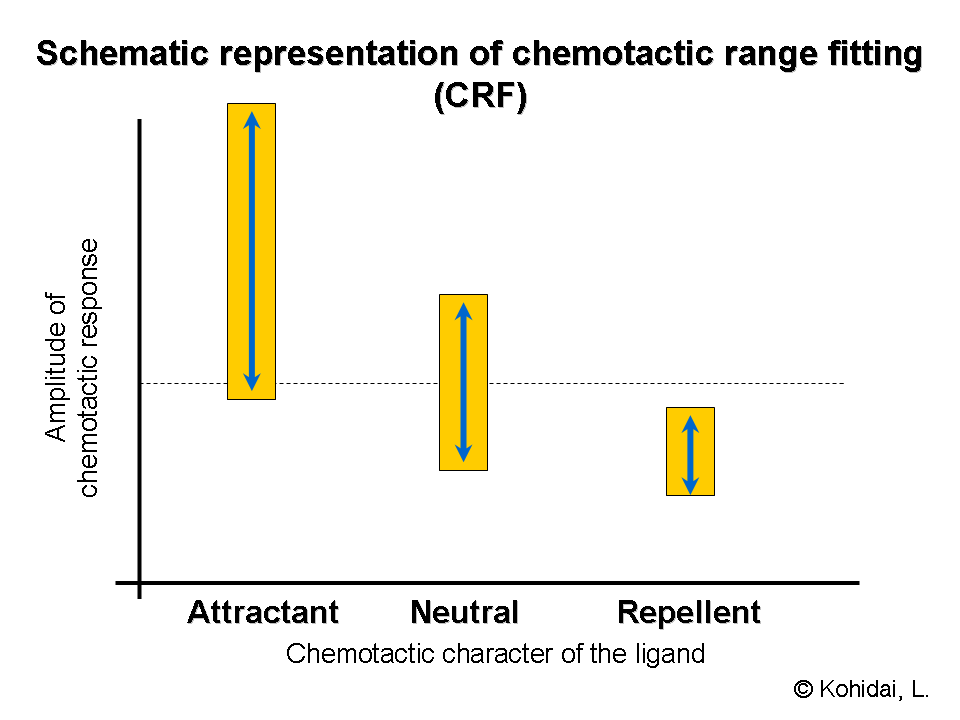
Chemotactic range fitting (CRF)
Chemotactic responses elicited by the ligand-receptor interactions are distinguished generally upon the optimal effective concentration(s) of the ligand. Nevertheless, correlation of the amplitude elicited and ratio of the responder cells compared to the total number are also characteristic features of the chemotactic signaling. Investigations of ligand families (e.g. amino acids or oligo peptides) proved that there is a fitting of ranges (amplitudes; number of responder cells) and chemotactic activities: chemoattractant moiety is accompanied by wide ranges, while chemorepellent character by narrow ranges.
Clinical significance
A changed migratory potential of cells has relatively high importance in the development of several clinical symptoms and syndromes. Altered chemotactic activity of extracellular (e.g. Escherichia coli) or intracellular (e.g. Listeria monocytogenes) pathogens itself represents a significant clinical target. Modification of endogenous chemotactic ability of these microorganisms by pharmaceutical agents can decrease or inhibit the ratio of infections or spreading of infectious diseases. Apart from infections, there are some other diseases where impaired chemotaxis is the primary ethiological factor, as in Chediak-Higashi syndrome where giant intracellular vesicles inhibit normal migration of cells.
| Type of disease | Chtx. increased | Chtx. decreased |
|---|---|---|
| Infections | inflammations | AIDS, Brucellosis |
| Chtx. results the disease | - | Chediak-Higashi syndrome, Kartagener syndrome |
| Chtx. is affected | atherosclerosis, arthritis, periodontitis, psoriasis, reperfusion injury, metastatic tumors | multiple sclerosis, Hodgkin disease, male infertility |
| Intoxications | asbestos, benzpyrene | salts of Hg and Cr, ozone (O3) |
Measurement of chemotaxis
A wide range of techniques is available to evaluate chemotactic activity of cells or the chemoattractant and chemorepellent character of ligands. The basic requirements of the measurement are as follows:
- concentration gradients can develop relatively fast and persist for a long time in the system
- chemotactic and chemokinetic activities are distinguished
- migration of cells is free towards and away on the axis of the concentration gradient
- detected responses are the results of active migration of cells
Despite the fact that an ideal chemotaxis assay is still not available, there are several protocols and pieces of equipment which offer good correspondence with the conditions described above. The most commonly used are:
- Agar plate assays
E.g. PP-chamber
- Two-chamber techniques
E.g. Boyden-chamber - Zigmond chamber - Dunn chambers - Multi-well chambers - Capillary techniques
- Others
E.g. T-maze technique - Opalescence technique - Orientation assays
(A more detailed chapter you can find under Chemotaxis assay)
For a cell to move, it requires a number of cellular components (such as cellular motors various enzymes, etc.), and for a cell to be able to move, it has to be able to change shape. Cell movement, broadly speaking can be of two types…
Hapoptatic (which means movement in response to physical and mechanical stimuli). Chemotactic (which is movement in response to a chemical gradient).
References
- ↑ Julius Adler and Wung-Wai Tso (1974). "Decision-Making in Bacteria: Chemotactic Response of Escherichia Coli to Conflicting Stimuli". Science. 184: 1292–4.
- ↑ http://research.microsoft.com/displayArticle.aspx?id=1572 retrieved November 6, 2006
- ↑ http://news.bbc.co.uk/2/hi/science/nature/6113522.stm retrieved November 6, 2006
- ↑ http://www.chemotaxis.usn.hu/CHTXhpg/CHTXresAA.htm
- ↑ Howard C. Berg (2003). "E. coli in motion". Springer-Verlag, NY. ISBN 0-387-00888-8.
- ↑ Kohidai L and Csaba G (1988). "Chemotaxis and chemotactic selection induced with cytokines (IL-8, RANTES and TNF alpha) in the unicellular Tetrahymena pyriformis". Cytokine. 10: 481–6.
External links
- Chemotaxis
- Cell Migration Gateway
- Global Existence for Chemotaxis with Finite Sampling Radius
- Bacterial Chemotaxis Depends on a Two-Component Signaling Pathway Activated by Histidine-Kinase-associated Receptors, Molecular Biology of the Cell 4th Edition © 2002 by Bruce Alberts, Alexander Johnson, Julian Lewis, Martin Raff, Keith Roberts, and Peter Walter.
- Figure 15-69. The two-component signaling pathway that enables chemotaxis receptors to control the flagellar motor during bacterial chemotaxis, Molecular Biology of the Cell 4th Edition © 2002 by Bruce Alberts, Alexander Johnson, Julian Lewis, Martin Raff, Keith Roberts, and Peter Walter.
Template:WH Template:WS Template:Jb1
ca:Quimiotaxi da:Kemotaksi de:Chemotaxis eo:Kemotaksiso it:Chemiotassi he:כימוטקסיס hu:Kemotaxis sv:Kemotaxis uk:Хемотаксис ur:کیمیحرک