Hepatitis D causes
|
Hepatitis D |
|
Diagnosis |
|
Treatment |
|
Hepatitis D causes On the Web |
|
American Roentgen Ray Society Images of Hepatitis D causes |
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [1]; Associate Editor(s)-in-Chief:
Overview
Taxonomy
Viruses; Deltavirus; Hepatitis delta virus
Biology
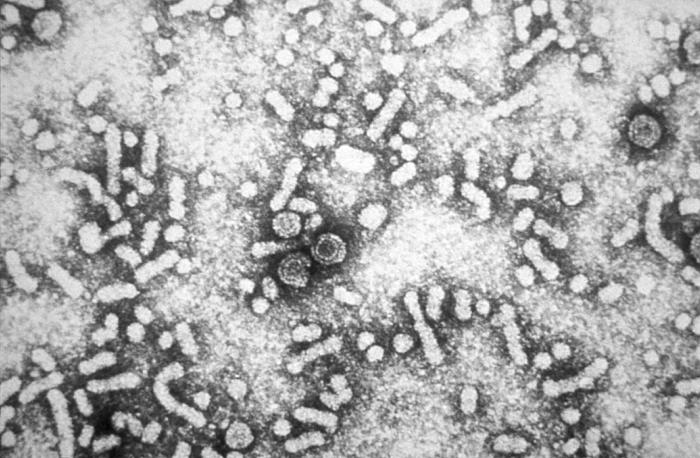 |
|
The HDV genome is a single, negative stranded, circular RNA molecule nearly 1.7 kb in length containing about 60% C+G. Hepatitis D virus(HDV) is the causative organism for Hepatitis D infection. HDV is only found in people who carry the hepatitis B virus. HDV may make a recent (acute) hepatitis B infection or an existing long-term (chronic) hepatitis B liver disease worse. It can even cause symptoms in people who carry hepatitis B virus but who never had symptoms. Hepatitis D infects about 15 million people worldwide. It occurs in 5% of people who carry hepatitis B. Risk factors include:
VirologyGenome structure and similarities to viroids The HDV genome exists as a negative sense, single-stranded, closed circular RNA. Because of a nucleotide sequence that is 70% self-complementary, the HDV genome forms a partially double stranded RNA structure that is described as rod-like.[2] With a genome of approximately 1700 nucleotides, HDV is the smallest "virus" known to infect animals. It has been proposed that HDV may have originated from a class of plant viruses called viroids.[3] Evidence in support of this hypothesis stems from the fact that both HDV and viroids exist as single-stranded, closed circular RNAs that have rod-like structures. Likewise, both HDV and viroids contain RNA sequences that can assume catalytically active structures called ribozymes. During viral replication, these catalytic RNAs are required in order to produce unit length copies of the genome from longer RNA concatamers. Finally, neither HDV nor viroids encode their own polymerase. Instead, replication of HDV and viroids requires a host polymerase that can utilize RNA as a template.[4] RNA polymerase II has been implicated as the polymerase responsible for the replication of HDV.[5][6] Normally RNA polymerase II utilizes DNA as a template and produces mRNA. Consequently, if HDV indeed utilizes RNA polymerase II during replication, it would be the only known pathogen capable of using a DNA-dependent polymerase as an RNA-dependent polymerase. Life CycleTo replicate efficiently, a virus requires the cooperation of the host cell at all stages of the replicative cycle:[7]
Synthesis of antigenomic RNA occurs in the nucleous, mediated by RNA polymerase I, whereas synthesis of genomic RNA takes place in the nucleoplasm, mediated by RNA Pol II.[8]
These elements are only assembled in the presence of the hepatitis B virus (helper virus). HBsAg and HDAg-L are necessary and sufficient for virus assembly, whereas HDV RNA or HDAg-S are not required, but are present, in viral particles. The primary initiation event for HDV assembly is the interaction of HDAg-L with HBsAg. HDAg is localized in the nuclei while and HBsAg is present in the cytoplasm of the infected cells. The mechanism of interaction between these two proteins remains unknown. That are different models that try to explain the mechanisms of viral RNA transcription and replication.[7][12][13] The Delta Antigens A significant difference between viroids and HDV is that, while viroids produce no proteins, HDV produces two proteins called the small and large delta antigens (HDAg-S and HDAg-L, respectively). These two proteins are produced from a single open reading frame. They are identical for 195 amino acids and differ only by the presence of an additional 19 amino acids at the C-terminus of HDAg-L. Despite having 90% identical sequences, these two proteins play diverging roles during the course of an infection. HDAg-S is produced in the early stages of an infection and is required for viral replication. HDAg-L, in contrast, is produced during the later stages of an infection, acts as an inhibitor of viral replication, and is required for assembly of viral particles. Evolution Three genotypes (I-III) were originally described. Genotype I has been isolated in Europe, North America, Africa and some Asia. Genotype II has been found in Japan, Taiwan, and Yakutia (Russia). Genotype III has been found exclusively in South America (Peru, Colombia, and Venezuela). Some genomes from Taiwan and the Okinawa islands have been difficult to type but have been placed in genotype 2. However it is not known that there are at least 8 genotypes of this virus (HDV-1 to HDV-8).[14]Phylogenetic studies suggest an African origin for this pathogen.[15] References
|