Genetic recombination
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [1]
Genetic recombination is the process by which a strand of DNA is broken and then joined to the end of a different DNA molecule. In eukaryotes recombination commonly occurs during meiosis as chromosomal crossover between paired chromosomes. This process leads to offspring having different combinations of genes from their parents and can produce new chimeric alleles. In evolutionary biology this shuffling of genes is thought to have many advantages, including that of allowing sexually reproducing organisms to avoid Muller's ratchet. However, a recombination pathway in DNA is any way by which a broken DNA molecule is reconnected to form a whole DNA strand.
In molecular biology "recombination" can also refer to artificial and deliberate recombination of disparate pieces of DNA, often from different organisms, creating what is called recombinant DNA.
Enzymes called recombinases catalyze natural recombination reactions. RecA, the recombinase found in E. coli, is responsible for the repair of DNA double strand breaks (DSBs). In yeast and other eukaryotic organisms there are two recombinases required for repairing DSBs. The RAD51 protein is required for mitotic and meiotic recombination and the DMC1 protein is specific to meiotic recombination.
Chromosomal crossover
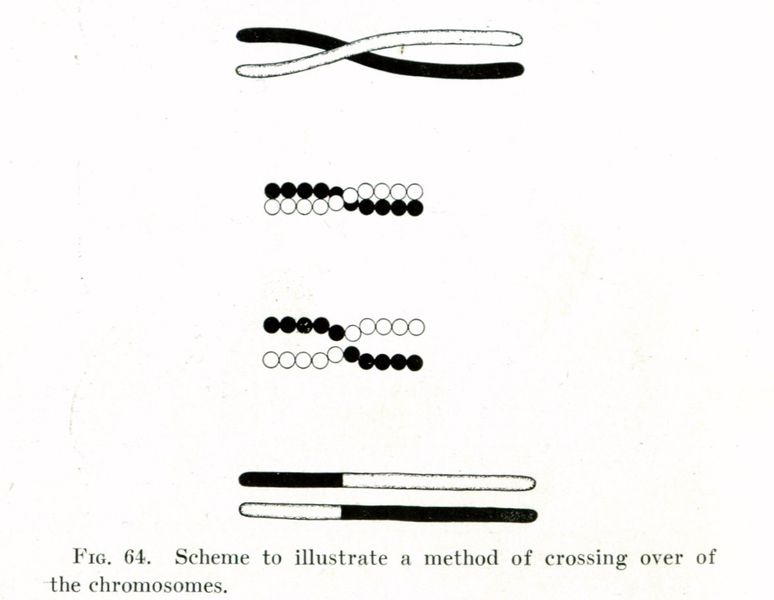
Chromosomal crossover refers to recombination between the paired chromosomes inherited from each of one's parents, generally occurring during meiosis. During prophase I the four available chromatids are in tight formation with one another. While in this formation, homologous sites on two chromatids can mesh with one another, and may exchange genetic information.
Because recombination can occur with small probability at any location along chromosome, the frequency of recombination between two locations depends on their distance. Chromosomes are expected to cross over at many points along their length; for genes sufficiently distant on the same chromosome the amount of crossover is so high that these genes effectively assort independently.
Conservative site-specific recombination
In conservative site-specific recombination, a mobile DNA element is inserted into a strand of DNA by means similar to that seen in crossover. A segment of DNA on the mobile element matches exactly with a segment of DNA on the target, allowing enzymes called integrases to insert the rest of the mobile element into the target. Integrases are a special type of Recombinases. Recombinases are enzymes which cleave the double stranded DNA at specific sites resulting in a loss of the Phosphodiester bonds. This reaction is stabilised by the formation of a covalent bond between the Recombinase and the DNA through a Phospho Tyrosine Bond.
Transpositional recombination
Another form of site-specific recombination, transpositional recombination does not require an identical strand of DNA in the mobile element to match with the target DNA. Instead, the integrases involved introduce nicks in both the mobile element and the target DNA, allowing the mobile DNA to enter the sequence. The nicks are then removed by ligases.
Nonhomologous recombination
Recombination between DNA sequences that contain no sequence homology, also referred to as nonhomologous end joining.
See also
External links
- The Holliday Model of Genetic Recombination
- Genetic+recombination at the US National Library of Medicine Medical Subject Headings (MeSH)
- Animated guide to homologous recombination.
References
- Alberts, B. et al., Molecular Biology of the Cell, 3rd Edition. Garland Publishing, 1994.
- Mayerhofer R, Koncz-Kalman Z, Nawrath C, Bakkeren G, Crameri A, Angelis K, Redei GP, Schell J, Hohn B, Koncz C. T-DNA integration: a mode of illegitimate recombination in plants. EMBO J. 1991 Mar;10(3):697-704.
Template:NCBI-scienceprimer Template:Genetic recombination
ar:تأشيب de:Rekombination he:שחלוף nl:Recombinatie (genetica) no:Rekombinasjon fi:Rekombinaatio sv:Rekombination (biologi) Template:Jb1