Nucleosome
|
WikiDoc Resources for Nucleosome |
|
Articles |
|---|
|
Most recent articles on Nucleosome |
|
Media |
|
Evidence Based Medicine |
|
Clinical Trials |
|
Ongoing Trials on Nucleosome at Clinical Trials.gov Clinical Trials on Nucleosome at Google
|
|
Guidelines / Policies / Govt |
|
US National Guidelines Clearinghouse on Nucleosome
|
|
Books |
|
News |
|
Commentary |
|
Definitions |
|
Patient Resources / Community |
|
Patient resources on Nucleosome Discussion groups on Nucleosome Patient Handouts on Nucleosome Directions to Hospitals Treating Nucleosome Risk calculators and risk factors for Nucleosome
|
|
Healthcare Provider Resources |
|
Causes & Risk Factors for Nucleosome |
|
Continuing Medical Education (CME) |
|
International |
|
|
|
Business |
|
Experimental / Informatics |
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [1]
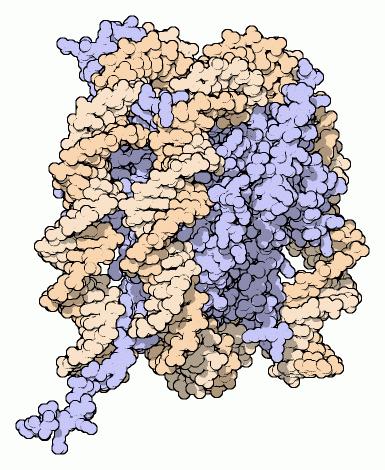
Nucleosomes form the fundamental repeating units of eukaryotic chromatin[1], which is used to pack the large eukaryotic genomes into the nucleus while still ensuring appropriate access to it (in mammalian cells approximately 2 m of linear DNA have to be packed into a nucleus of roughly 10 µm diameter). Nucleosomes are folded through a series of successively higher order structures to eventually form a chromosome; this both compacts DNA and creates an added layer of regulatory control which ensures correct gene expression. Nucleosomes are thought to carry epigenetically inherited information in the form of covalent modifications of their core histones. The nucleosome hypothesis proposed by Don and Ada Olins[2] and Roger Kornberg[3][4] in 1974, was a paradigm shift for understanding eukaryotic gene expression.
The nucleosome core particle consists of approximately 147[5] base pairs of DNA wrapped in 1.67 left-handed superhelical turns around a histone octamer consisting of 2 copies each of the core histones H2A, H2B, H3, and H4.[6] Linker histones such as H1 and its isoforms are involved in chromatin compaction and sit at the base of the nucleosome near the DNA entry and exit binding to the linker region of the DNA.[7] Non-condensed nucleosomes without the linker histone resemble "beads on a string of DNA" under an electron microscope.[8]
In contrast to most eukaryotic cells mature sperm cells largely use protamines to package their genomic DNA, most likely to achieve an even higher packaging ratio.[9] Histone equivalents and a simplified chromatin structure have also been found in Archea[10], proving that eukaryotes are not the only organisms that use nucleosomes.
Structure
Structure of the core particle
Overview
Early structural studies provided evidence that an octamer of histone proteins wraps DNA around itself in about two turns of a left-handed superhelix. In 1997 the first near atomic resolution crystal structure of the nucleosome was solved by the Richmond group showing some of the most important details of the particle. The structure of over 20 different nucleosome core particles have been solved to date[11], including those containing histone variants and histones from different species. The structure of the nucleosome core particle is remarkably conserved, and even a change of over 100 residues between frog and yeast histones results in electron density maps with an overall root mean square deviation (r.m.s.d) of only 1.6Å[12].
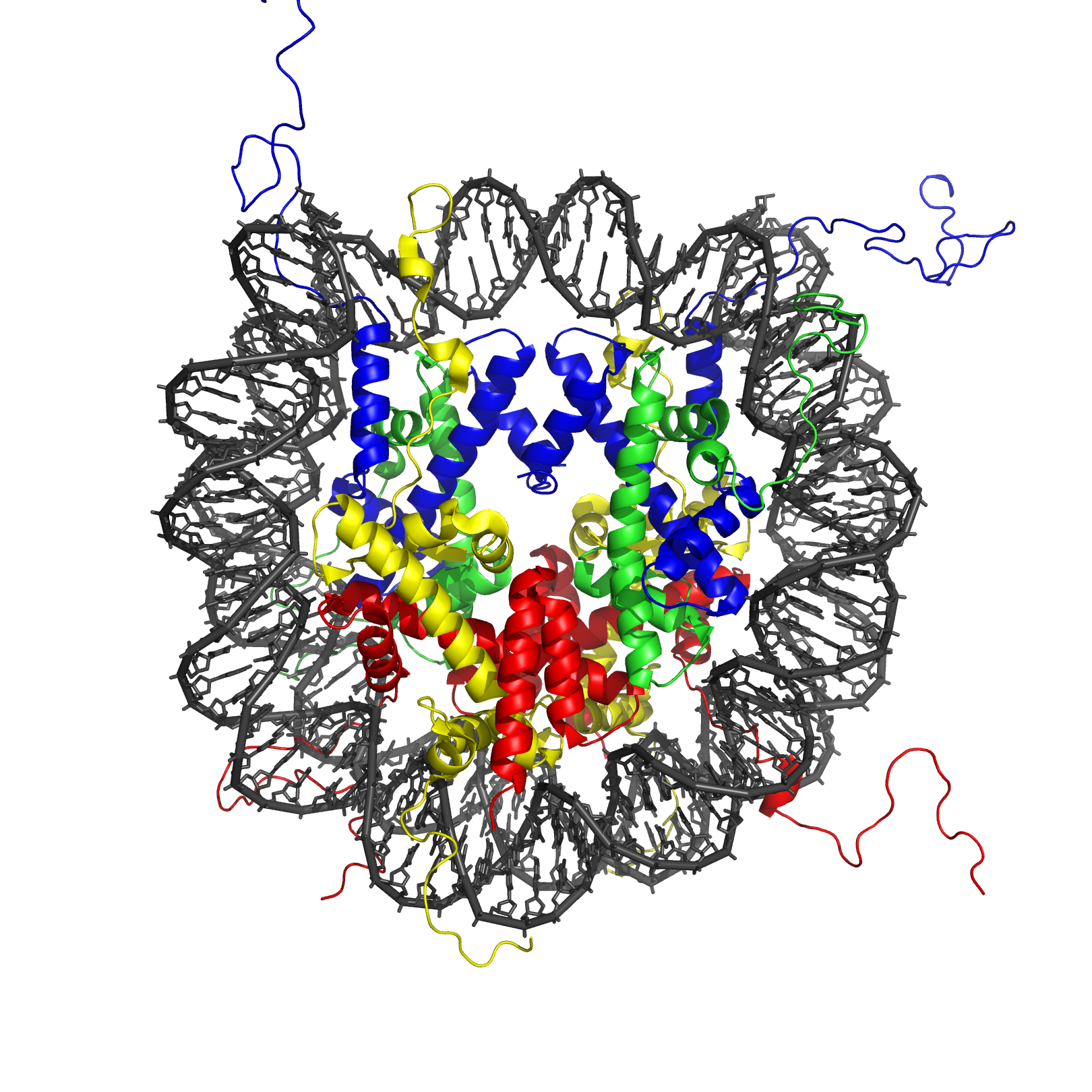
The nucleosome core particle
The nucleosome core particle (shown in the figure) consists of about 146[5] bp of DNA wrapped in 1.67 left-handed superhelical turns around the histone octamer, consisting of 2 copies each of the core histones H2A, H2B, H3, and H4. Adjacent nucleosomes are joined by a stretch of free DNA termed "linker DNA" which varies from 10 - 80 bp in length depending on species and tissue type[10]).
Protein interactions within the nucleosome
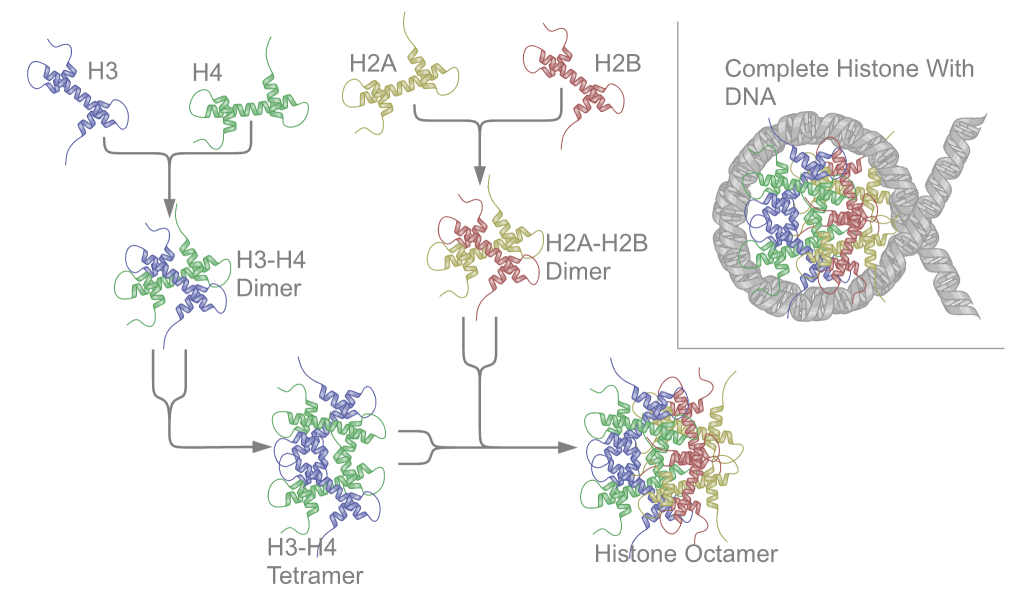
The core histone proteins contain a characteristic structural motif termed the "histone fold" which consists of three alpha-helices (α1-3) separated by two loops (L1-2). In solution the histones form H2A-H2B heterodimers and H3-H4 heterotetramers. Histones dimerise about their long α2 helices in an anti-parallel orientation, and in the case of H3 and H4, two such dimers form a 4-helix bundle stabilised by extensive H3-H3’ interaction. The H2A/H2B dimer binds onto the H3/H4 tetramer due to interactions between H4 and H2B which include the formation of a hydrophobic cluster[6]. The histone octamer is formed by a central H3/H4 tetramer sandwiched between two H2A/H2B dimers. Due to the highly basic charge of all four core histones, the histone octamer is only stable in the presence of DNA or very high salt concentrations.
Histone - DNA interactions
The nucleosome contains over 120 direct protein-DNA interactions and several hundred water mediated ones[13]. Direct protein - DNA interactions are not spread evenly about the octamer surface but rather located at discrete sites. These are due to the formation of two types of DNA binding sites within the octamer; the α1α1 site which uses the α1 helix from two adjacent histones and the L1L2 site formed by the L1 and L2 loops. Salt links and hydrogen bonding between both side chain basic and hydroxyl groups and main chain amides with the DNA backbone phosphates form the bulk of interactions with the DNA. This is important given that the ubiquitous distribution of nucleosomes along genomes requires it to be a non-sequence-specific DNA-binding factor. Although nucleosomes tend to prefer some DNA sequences over others[14], they are capable of binding practically to any sequence, which is thought to be due to the flexibility in the formation of these water-mediated interactions. In addition, non-polar interactions are made between protein side chains and the deoxyribose groups, and an arginine side chain intercalates into the DNA minor groove at all 14 sites it faces the octamer surface. The distribution and strength of DNA binding sites about the octamer surface distorts the DNA within the nucleosome core. The DNA is non-uniformly bent and also contains twist defects. The twist of free B-form DNA in solution is 10.5 bp per turn, however, the overall twist of nucleosomal DNA is only 10.2 bp per turn, varying from a value of 9.4 to 10.9 bp per turn.
Histone tail domains
The histone tail extensions constitute up to 30% by mass of histones, but are not visible in the crystal structures of nucleosomes due to their high intrinsic flexibility and have been thought to be largely unstructured[15]. The N-terminal tails of histones H3 and H2B pass through a channel formed by the minor grooves of the two DNA strands, protruding from the DNA every 20 bp. The N-terminal tail of histone H4 on the other hand has a region of highly basic amino acids (16-25) which, in the crystal structure, forms an interaction with the highly acidic surface region of a H2A-H2B dimer of another nucleosome, being potentially relevant for the higher-order structure of nucleosomes. This interaction is thought to occur also under physiological conditions and suggests that acetylation of the H4 tail distorts the higher order structure of chromatin.
Higher order structure
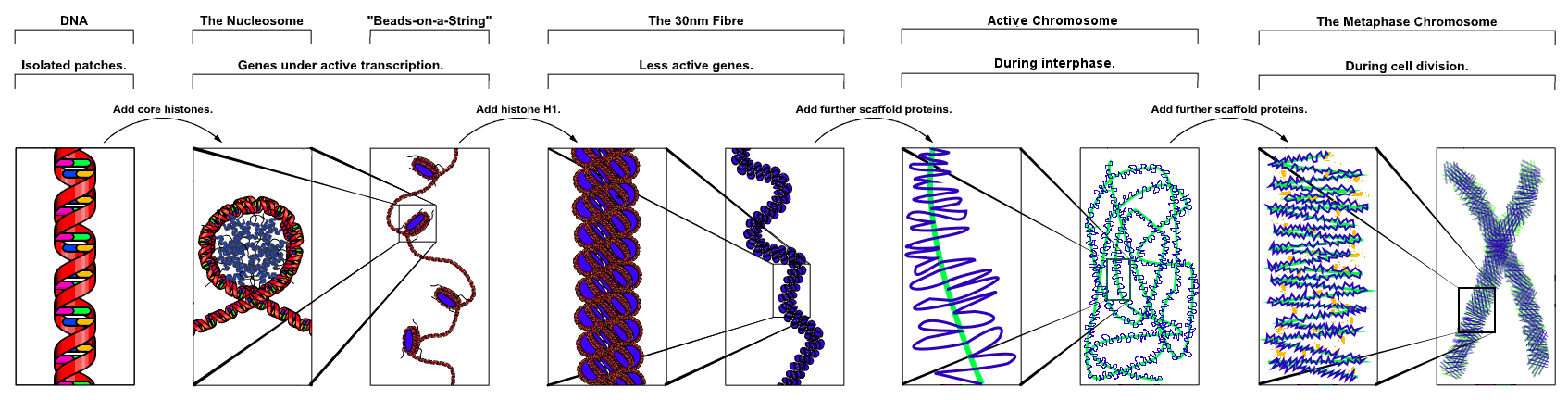

The organization of the DNA that is achieved by the nucleosome can not fully explain the packaging of DNA observed in the cell nucleus. Further compaction of chromatin into the cell nucleus is necessary, but is not yet well understood. The current understanding[16] is that repeating nucleosomes with intervening "linker" DNA form a 10-nm-fiber, known descriptively as "beads on a string", and have a packing ratio of about five to ten[10]. A chain of nucleosomes can be arranged in a 30 nm fiber, a compacted structure with a packing ratio of ~50[10] and whose formation is dependent on the presence of the H1 histone.
A crystal structure of a tetranucleosome has been presented and used to build up a proposed structure of the 30 nm fiber as a two-start helix.[17] There is still a certain amount of contention regarding this model as it is incompatible with recent electron microscopy data[18]. Beyond this, the structure of chromatin is poorly understood, but it is classically suggested that the 30 nm fiber is arranged into loops along a central protein scaffold to form transcriptionally active euchromatin. Further compaction leads to transcriptionally inactive heterochromatin.
Nucleosome dynamics
Although the nucleosome is a very stable protein-DNA complex, it is not static and has been shown to undergo a number of different structural re-arrangements including nucleosome sliding and DNA site exposure.
Nucleosome sliding
Work performed in the Bradbury laboratory showed that nucleosomes reconstituted onto the 5S DNA positioning sequence were able to reposition themselves translationally onto adjacent sequences when incubated thermally[19]. Later work showed that this repositioning did not require disruption of the histone octamer but was consistent with nucleosomes being able to “slide” along the DNA in cis. In 2008, It was further revealed that CTCF binding sites act as nucleosome positioning anchors so that, when used to align various genomic signals, multiple flanking nucleosomes can be readily identified[20]. Although nucleosomes are intrinsically mobile, eukaryotes have evolved a large family of ATP-dependent chromatin remodelling enzymes to alter chromatin structure, many of which do so via nucleosome sliding.
DNA site exposure
Work from the Widom laboratory has shown that nucleosomal DNA is in equilibrium between a wrapped and unwrapped state. Measurements of these rates using time resolved FRET revealed that DNA within the nucleosome remains fully wrapped for only 250ms before it is unwrapped for 10-50ms and then rapidly rewrapped [21]. This implies that DNA does not need to be actively dissociated from the nucleosome but that there is a significant fraction of time during which it is fully accessible. Indeed, this can be extended to the observation that introducing a DNA binding sequence within the nucleosome increases the accessibility of adjacent regions of DNA when bound [22]. This propensity for DNA within the nucleosome to “breathe” is predicted to have important functional consequences for all DNA binding proteins that operate in a chromatin environment.
Modulating nucleosome structure
Eukaryotic genomes are ubiquitously associated into chromatin; however, cells need to spatially and temporally regulate specific loci independently of bulk chromatin. In order to achieve the high level of control required to co-ordinate nuclear processes such as DNA replication, repair and transcription, cells have developed a variety of means to locally and specifically modulate chromatin structure and function. This can involve covalent modification of histones, the incorporation of histone variants and non-covalent remodelling by ATP-dependent remodelling enzymes.
Histone post-translational modifications
Since they were discovered in the mid 1960’s histone modifications have been predicted to affect transcription[23]. The fact that most of the early post-translational modifications found were concentrated within the tail extensions that protrude from the nucleosome core lead to two main theories regarding the mechanism of histone modification. The first of the theories suggested that they may affect electrostatic interactions between the histone tails and DNA to “loosen” chromatin structure. Later it was proposed that combinations of these modifications may create binding epitopes with which to recruit other proteins[24]. Recently, given that more modifications have been found in the structured regions of histones it has been put forward that these modifications may affect histone-DNA[25] and histone-histone[26] interactions within the nucleosome core. Some modifications have been shown to be correlated with gene silencing, others seem to be correlated with gene activation. Common modifications include acetylation, methylation or ubiquitination of lysine; methylation of arginine and phosphorylation of serine. The information stored in this way is considered epigenetic since it is not encoded in the DNA but is still inherited to daughter cells. The maintenance of a repressed or activated status of a gene is often necessary for cellular differentiation.[10]
Histone variants
Whilst histones are remarkably conserved throughout evolution, several variant forms have been identified. Interestingly, this diversification of histone function is restricted to H2A and H3, with H2B and H4 being mostly invariant. H2A can be replaced by H2AZ (which leads to reduced nucleosome stability) or H2AX (which is associated with DNA repair and T cell differentiation) whereas the inactive X chromosomes in mammals are enriched in macroH2A. H3 can be replaced by H3.3 (which correlates with activate genes) and in centromeres H3 is replaced by CENPA.[10]
ATP-dependent nucleosome remodelling
A number of distinct reactions are associated with the term ATP-dependent chromatin remodelling. Remodelling enzymes have been shown to slide nucleosomes along DNA[27],disrupt histone-DNA contacts to the extent of destabilising the H2A/H2B dimer[28][29] and to generate negative superhelical torsion in DNA and chromatin[30]. Recently, the Swr1 remodelling enzyme has been shown to introduce the variant histone H2A.Z into nucleosomes[31]. At present, it is not clear if all of these represent distinct reactions or merely alternative outcomes of a common mechanism. What is shared between all, and indeed the hallmark of ATP-dependent chromatin remodelling, is that they all result in altered DNA accessibility. Studies looking at gene activation in vivo[32] and, more astonishingly, remodelling in vitro[33] has revealed that chromatin remodelling events and transcription-factor binding are cyclical and periodic in nature. While the consequences of this for the reaction mechanism of chromatin remodelling are not known, the dynamic nature of the system may allow it to respond faster to external stimuli.
Dynamic nucleosome remodelling across the Yeast genome
Studies in 2007 have catalogued nucleosome positions in yeast and shown that nucleosomes are enriched in promoter regions [34][35][36]. About 80% of the yeast genome appears to be covered by nucleosomes and the pattern of nucleosome positioning clearly relates to DNA regions that regulate transcription and regions that are transcribed. Most recently, a new study examined ‘’dynamic changes’’ in nucleosome repositioning during a global transcriptional reprogramming event to elucidate the effects on nucleosome displacement during genome-wide transcriptional changes in yeast (Saccharomyces cerevisiae) [37]. The results suggested that nucleosomes that were localized to promoter regions are displaced in response to stress (like heat shock). In addition, the removal of nucleosomes usually corresponded to transcriptional activation and the replacement of nucleosomes usually corresponded to transcriptional repression, presumably because transcription factor binding sites became more or less accessible, respectively. In general, only one or two nucleosomes were repositioned at the promoter to effect these transcriptional changes. However, even in chromosomal regions that were not associated with transcriptional changes, nucleosome repositioning was observed, suggesting that the covering and uncovering of transcriptional DNA does not necessarily produce a transcriptional event.
Nucleosome assembly in vitro
Nucleosomes can be assembled in vitro by either using purified native or recombinant histones.[38][39] One standard technique of loading the DNA around the histones involves the use of salt dialysis. A reaction consisting of the histone octamers and a naked DNA template can be incubated together at a salt concentration of 2 M. By steadily decreasing the salt concentration, the DNA will equilibrate to a position where it is wrapped around the histone octamers, forming nucleosomes. In appropriate conditions, this reconstitution process allows for the nucleosome positioning affinity of a given sequence to be mapped experimentally. [40]
References
- ↑ Alberts, B., et al. Molecular Biology of the Cell, Fourth Ed., 2002, p. 207
- ↑ Olins AL and Olins DE, "Spheroid Chromatin Units (nu Bodies)", Science (1974); 183: 330 - 332
- ↑ McDonald D, "Milestone 9, (1973-1974) The nucleosome hypothesis: An alternative string theory", Nature Milestones: Gene Expression. (2005) Dec 1; http://www.nature.com/milestones/geneexpression/milestones/articles/milegene09.html
- ↑ Kornberg, RD, "Chromatin structure: a repeating unit of histones and DNA", Science. (1974); 184: 868–871
- ↑ 5.0 5.1 In different crystals, values of 146 and 147 basepairs were observed
- ↑ 6.0 6.1 Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ, "Crystal Structure of the Nucleosome Core Particle at 2.8 Å Resolution", Nature. 1997 Sep 18; 389 (6648): 251-60.
- ↑ Position and orientation of the globular domain of linker histone H5 on the nucleosome : Abstract : Nature
- ↑ Involvement of histone H1 in the organization of the nucleosome and of the salt-dependent superstructures of chromatin - Thoma et al. 83 (2): 403 - The Journal of Cell Biology
- ↑ Clark, H.J. Nuclear and chromatin composition of mammalian gametes and early embryos. Biochem Cell Biol. 1992 Oct-Nov;70(10-11):856-66, PMID 1297351
- ↑ 10.0 10.1 10.2 10.3 10.4 10.5 Felsenfeld G, Groudine M (2003). "Controlling the double helix". Nature. 421 (6921): 448–53. doi:10.1038/nature01411. PMID 12540921. Unknown parameter
|month=ignored (help) - ↑ Chakravarthy S, Park YJ, Chodaparambil J, Edayathumangalam RS, Luger K (2005). "Structure and dynamic properties of nucleosome core particles". FEBS Lett. 579 (4): 895–8. doi:10.1016/j.febslet.2004.11.030. PMID 15680970. Unknown parameter
|month=ignored (help) - ↑ White CL, Suto RK, Luger K (2001). "Structure of the yeast nucleosome core particle reveals fundamental changes in internucleosome interactions". EMBO J. 20 (18): 5207–18. doi:10.1093/emboj/20.18.5207. PMC 125637. PMID 11566884. Unknown parameter
|month=ignored (help) - ↑ Davey CA, Sargent DF, Luger K, Maeder AW, Richmond TJ (2002). "Solvent mediated interactions in the structure of the nucleosome core particle at 1.9 a resolution". J. Mol. Biol. 319 (5): 1097–113. doi:10.1016/S0022-2836(02)00386-8. PMID 12079350. Unknown parameter
|month=ignored (help) - ↑ Segal E. et al., "A genomic code for nucleosome positioning", Nature 442, 772-778 (17 August 2006)
- ↑ Zheng C, Hayes JJ (2003). "Structures and interactions of the core histone tail domains". Biopolymers. 68 (4): 539–46. doi:10.1002/bip.10303. PMID 12666178. Unknown parameter
|month=ignored (help) - ↑ Chakravarthy S, Park YJ, Chodaparambil J, Edayathumangalam RS, Luger K, "Structure and dynamic properties of nucleosome core particles", FEBS Letters. (2005) Feb 7; 579 (4): 895-898.
- ↑ Schalch T, Duda S, Sargent DF, Richmond TJ, "X-ray structure of a tetranucleosome and its implications for the chromatin fibre", Nature. (2005) Jul 7; 436: 138-141.
- ↑ Robinson PJ, Fairall L, Huynh VA, Rhodes D (2006). "EM measurements define the dimensions of the "30-nm" chromatin fiber: evidence for a compact, interdigitated structure". Proc. Natl. Acad. Sci. U.S.A. 103 (17): 6506–11. doi:10.1073/pnas.0601212103. PMC 1436021. PMID 16617109. Unknown parameter
|month=ignored (help) - ↑ Pennings S, Muyldermans S, Meersseman G, Wyns L (1989). "Formation, stability and core histone positioning of nucleosomes reassembled on bent and other nucleosome-derived DNA". J. Mol. Biol. 207 (1): 183–92. PMID 2738923. Unknown parameter
|month=ignored (help) - ↑ Fu Y, Sinha M, Peterson CL, Weng Z (2008). "The insulator binding protein CTCF positions 20 nucleosomes around its binding sites across the human genome". PLoS genetics. 4 (7): e1000138. doi:10.1371/journal.pgen.1000138. PMC 2453330. PMID 18654629.
- ↑ Li G, Levitus M, Bustamante C, Widom J (2005). "Rapid spontaneous accessibility of nucleosomal DNA". Nat. Struct. Mol. Biol. 12 (1): 46–53. doi:10.1038/nsmb869. PMID 15580276. Unknown parameter
|month=ignored (help) - ↑ Li G, Widom J (2004). "Nucleosomes facilitate their own invasion". Nat. Struct. Mol. Biol. 11 (8): 763–9. doi:10.1038/nsmb801. PMID 15258568. Unknown parameter
|month=ignored (help) - ↑ ALLFREY VG, FAULKNER R, MIRSKY AE (1964). "ACETYLATION AND METHYLATION OF HISTONES AND THEIR POSSIBLE ROLE IN THE REGULATION OF RNA SYNTHESIS". Proc. Natl. Acad. Sci. U.S.A. 51: 786–94. PMC 300163. PMID 14172992. Unknown parameter
|month=ignored (help) - ↑ Strahl BD, Allis CD (2000). "The language of covalent histone modifications". Nature. 403 (6765): 41–5. doi:10.1038/47412. PMID 10638745. Unknown parameter
|month=ignored (help) - ↑ Cosgrove MS, Boeke JD, Wolberger C (2004). "Regulated nucleosome mobility and the histone code". Nat. Struct. Mol. Biol. 11 (11): 1037–43. doi:10.1038/nsmb851. PMID 15523479. Unknown parameter
|month=ignored (help) - ↑ Ye J, Ai X, Eugeni EE; et al. (2005). "Histone H4 lysine 91 acetylation a core domain modification associated with chromatin assembly". Mol. Cell. 18 (1): 123–30. doi:10.1016/j.molcel.2005.02.031. PMID 15808514. Unknown parameter
|month=ignored (help) - ↑ Whitehouse I, Flaus A, Cairns BR, White MF, Workman JL, Owen-Hughes T (1999). "Nucleosome mobilization catalysed by the yeast SWI/SNF complex". Nature. 400 (6746): 784–7. doi:10.1038/23506. PMID 10466730. Unknown parameter
|month=ignored (help) - ↑ Kassabov SR, Zhang B, Persinger J, Bartholomew B (2003). "SWI/SNF unwraps, slides, and rewraps the nucleosome". Mol. Cell. 11 (2): 391–403. PMID 12620227. Unknown parameter
|month=ignored (help) - ↑ Bruno M, Flaus A, Stockdale C, Rencurel C, Ferreira H, Owen-Hughes T (2003). "Histone H2A/H2B dimer exchange by ATP-dependent chromatin remodeling activities". Mol. Cell. 12 (6): 1599–606. PMID 14690611. Unknown parameter
|month=ignored (help) - ↑ Havas K, Flaus A, Phelan M; et al. (2000). "Generation of superhelical torsion by ATP-dependent chromatin remodeling activities". Cell. 103 (7): 1133–42. PMID 11163188. Unknown parameter
|month=ignored (help) - ↑ Mizuguchi G, Shen X, Landry J, Wu WH, Sen S, Wu C (2004). "ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex". Science (journal). 303 (5656): 343–8. doi:10.1126/science.1090701. PMID 14645854. Unknown parameter
|month=ignored (help) - ↑ Métivier R, Penot G, Hübner MR; et al. (2003). "Estrogen receptor-alpha directs ordered, cyclical, and combinatorial recruitment of cofactors on a natural target promoter". Cell. 115 (6): 751–63. PMID 14675539. Unknown parameter
|month=ignored (help) - ↑ Nagaich AK, Walker DA, Wolford R, Hager GL (2004). "Rapid periodic binding and displacement of the glucocorticoid receptor during chromatin remodeling". Mol. Cell. 14 (2): 163–74. PMID 15099516. Unknown parameter
|month=ignored (help) - ↑ Albert I, Mavrich TN, Tomsho LP, Qi J, Zanton SJ; et al. (2007). "Translational and rotational settings of H2A.Z nucleosomes across the Saccharomyces cerevisiae genome". Nature. 446: 572–576. doi:10.1038/nature05632. PMID 17392789.
- ↑ Li B, Carey M, Workman JL (2007). "The Role of Chromatin during Transcription". Cell. 128: 707–719. doi:10.1016/j.cell.2007.01.015. PMID 17320508.
- ↑ Whitehouse I, Rando OJ, Delrow J, Tsukiyama T (2007). "Chromatin remodelling at promoters suppresses antisense transcription". Nature. 450: 1031–1035. doi:10.1038/nature06391. PMID 18075583.
- ↑ Shivaswamy S, Bhinge A, Zhao Y, Jones S, Hirst M; et al. (2007). "Dynamic Remodeling of Individual Nucleosomes Across a Eukaryotic Genome in Response to Transcriptional Perturbation". PLoS Biol. 6 (3): e65. doi:10.1371/journal.pbio.0060065. PMID 18351804.
- ↑ Hayes, JJ, Lee, K.-M. In vitro reconstitution and analysis of mononucleosomes containing defined DNAs and proteins, Methods (1997), 12: 2-9, PMID 9169189.
- ↑ Dyer PN, Edayathumangalam RS, White CL, Bao Y, Chakravarthy S, Muthurajan UM, Luger K, "Reconstitution of nucleosome core particles from recombinant histones and DNA", Methods in Enzymology (2004); 375: 23-44.
- ↑ Yenidunya A, Davey C, Clark D, Felsenfeld G, Allan J. "Nucleosome positioning on chicken and human globin gene promoters in vitro. Novel mapping techniques.", Journal of Molecular Biology (1994); 237: 401-14.
External links
ca:Nucleosoma de:Nukleosom fa:نوکلئوزوم ko:뉴클레오좀 it:Nucleosoma he:נוקלאוזום lt:Nukleosoma nl:Nucleosoom oc:Nucleosòma sl:Nukleosom sr:Нуклеозом fi:Nukleosomi sv:Nukleosom uk:Нуклеосома